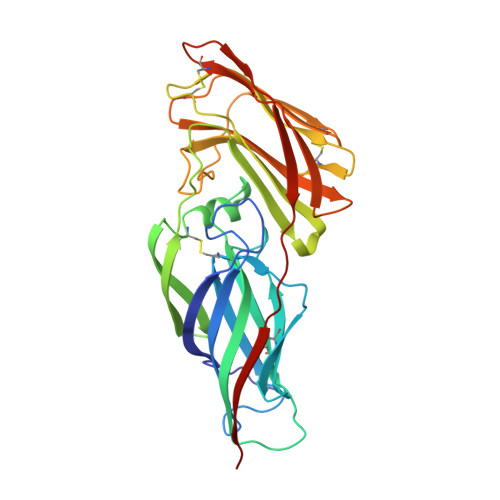
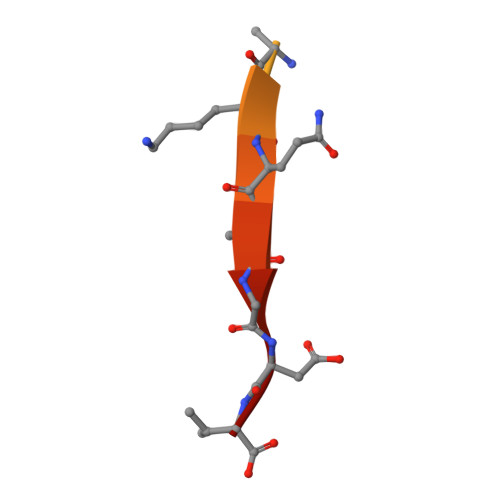
Structural Basis for the Broad Specificity to Host- Cell Ligands by the Pathogenic Fungus Candida Albicans.
Salgado, P.S., Yan, R., Taylor, J.D., Burchell, L., Jones, R., Hoyer, L.L., Matthews, S.J., Simpson, P.J., Cota, E.(2011) Proc Natl Acad Sci U S A 108: 15775
- PubMed: 21896717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1103496108
- Primary Citation Related Structures:
2Y7L, 2Y7M, 2Y7N, 2Y7O, 2YLH - PubMed Abstract:
Candida albicans is the most prevalent fungal pathogen in humans and a major source of life-threatening nosocomial infections. The Als (agglutinin-like sequence) glycoproteins are an important virulence factor for this fungus and have been associated with binding of host-cell surface proteins and small peptides of random sequence, the formation of biofilms and amyloid fibers. High-resolution structures of N-terminal Als adhesins (NT-Als; up to 314 amino acids) show that ligand recognition relies on a motif capable of binding flexible C termini of peptides in extended conformation. Central to this mechanism is an invariant lysine that recognizes the C-terminal carboxylate of ligands at the end of a deep-binding cavity. In addition to several protein-peptide interactions, a network of water molecules runs parallel to one side of the ligand and contributes to the recognition of diverse peptide sequences. These data establish NT-Als adhesins as a separate family of peptide-binding proteins and an unexpected adhesion system for primary, widespread protein-protein interactions at the Candida/host-cell interface.
- Division of Molecular Biosciences, Imperial College London, Exhibition Road, South Kensington SW7 2AZ, United Kingdom.
Organizational Affiliation: