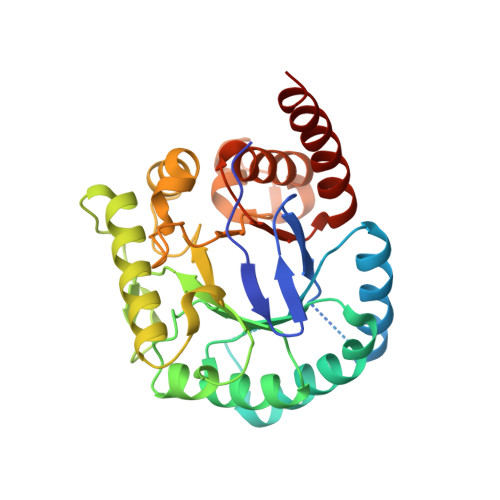
Crystal Structures of Burkholderia Cenocepacia Dihydropteroate Synthase in the Apo-Form and Complexed with the Product 7,8-Dihydropteroate.
Morgan, R.E., Batot, G.O., Dement, J.M., Rao, V.A., Eadsforth, T.C., Hunter, W.N.(2011) BMC Struct Biol 11: 21
- PubMed: 21554707 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-11-21
- Primary Citation Related Structures:
2Y5J, 2Y5S - PubMed Abstract:
The enzyme dihydropteroate synthase (DHPS) participates in the de novo synthesis of folate cofactors by catalyzing the formation of 7,8-dihydropteroate from condensation of p-aminobenzoic acid with 6-hydroxymethyl-7,8-dihydropteroate pyrophosphate. DHPS is absent from humans, who acquire folates from diet, and has been validated as an antimicrobial therapeutic target by chemical and genetic means. The bacterium Burkholderia cenocepacia is an opportunistic pathogen and an infective agent of cystic fibrosis patients. The organism is highly resistant to antibiotics and there is a recognized need for the identification of new drugs against Burkholderia and related Gram-negative pathogens. Our characterization of the DHPS active site and interactions with the enzyme product are designed to underpin early stage drug discovery. An efficient recombinant protein expression system for DHPS from B. cenocepacia (BcDHPS) was prepared, the dimeric enzyme purified in high yield and crystallized. The structure of the apo-enzyme and the complex with the product 7,8-dihydropteroate have been determined to 2.35 Å and 1.95 Å resolution respectively in distinct orthorhombic crystal forms. The latter represents the first crystal structure of the DHPS-pterin product complex, reveals key interactions involved in ligand binding, and reinforces data generated by other structural studies. Comparisons with orthologues identify plasticity near the substrate-binding pocket and in particular a range of loop conformations that contribute to the architecture of the DHPS active site. These structural data provide a foundation for hit discovery. An intriguing observation, an artifact of the analysis, that of a potential sulfenamide bond within the ligand complex structure is mentioned. Structural similarities between BcDHPS and orthologues from other Gram-negative species are evident as expected on the basis of a high level of sequence identity. The presence of 7,8-dihydropteroate in the binding site provides details about ligand recognition by the enzyme and the different states of the enzyme allow us to visualize distinct conformational states of loops adjacent to the active site. Improved drugs to combat infections by Burkholderia sp. and related Gram-negative bacteria are sought and our study now provides templates to assist that process and allow us to discuss new ways of inhibiting DHPS.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dow Street, Dundee, DD1 5EH, UK.
Organizational Affiliation: