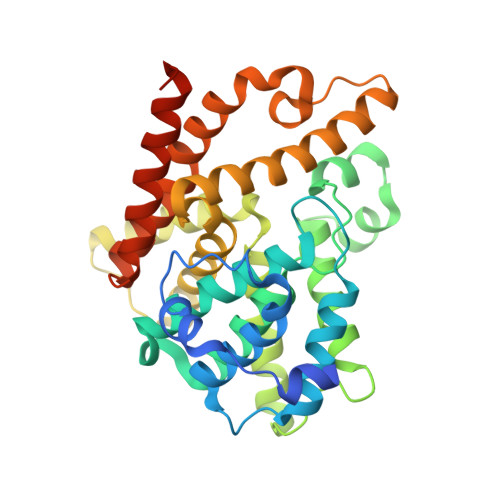
Triazoloquinazolines as a Novel Class of Phosphodiesterase 10A (Pde10A) Inhibitors.
Kehler, J., Ritzen, A., Langgard, M., Petersen, S.L., Farah, M.M., Bundgaard, C., Christoffersen, C.T., Nielsen, J., Kilburn, J.P.(2011) Bioorg Med Chem Lett 21: 3738
- PubMed: 21602043 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2011.04.067
- Primary Citation Related Structures:
2Y0J - PubMed Abstract:
Novel triazoloquinazolines have been found as phosphodiesterase 10A (PDE10A) inhibitors. Structure-activity studies improved the initial micromolar potency which was found in the lead compound by a 100-fold identifying 5-(1H-benzoimidazol-2-ylmethylsulfanyl)-2-methyl-[1,2,4]triazolo[1,5-c]quinazoline, 42 (PDE10A IC(50)=12 nM) as the most potent compound from the series. Two X-ray structures revealed novel binding modes to the catalytic site of the PDE10A enzyme.
- H. Lundbeck A/S, Department of Discovery Chemistry & DMPK, Valby, Denmark. jke@lundbeck.com
Organizational Affiliation: