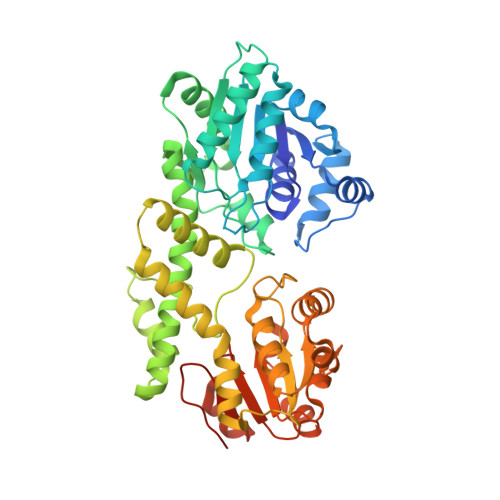
Structure of Burkholderia Cepacia Udp-Glucose Dehydrogenase (Ugd) Bcec and Role of Tyr10 in Final Hydrolysis of Ugd Thioester Intermediate.
Rocha, J., Popescu, A.O., Borges, P., Mil-Homens, D., Moreira, L.M., Sa-Correia, I., Fialho, A.M., Frazao, C.(2011) J Bacteriol 193: 3978
- PubMed: 21602353 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.01076-10
- Primary Citation Related Structures:
2Y0C, 2Y0D, 2Y0E - PubMed Abstract:
Members of the Burkholderia cepacia complex (BCC) are serious respiratory pathogens in immunocompromised individuals and in patients with cystic fibrosis (CF). They are exceptionally resistant to many antimicrobial agents and have the capacity to spread between patients, leading to a decline in lung function and necrotizing pneumonia. BCC members often express a mucoid phenotype associated with the secretion of the exopolysaccharide (EPS) cepacian. There is much evidence supporting the fact that cepacian is a major virulence factor of BCC. UDP-glucose dehydrogenase (UGD) is responsible for the NAD-dependent 2-fold oxidation of UDP-glucose (UDP-Glc) to UDP-glucuronic acid (UDP-GlcA), which is a key step in cepacian biosynthesis. Here, we report the structure of BceC, determined at 1.75-Å resolution. Mutagenic studies were performed on the active sites of UGDs, and together with the crystallographic structures, they elucidate the molecular mechanism of this family of sugar nucleotide-modifying enzymes. Superposition with the structures of human and other bacterial UGDs showed an active site with high structural homology. This family contains a strictly conserved tyrosine residue (Y10 in BceC; shown in italics) within the glycine-rich motif (GXGYXG) of its N-terminal Rossmann-like domain. We constructed several BceC Y10 mutants, revealing only residual dehydrogenase activity and thus highlighting the importance of this conserved residue in the catalytic activity of BceC. Based on the literature of the UGD/GMD nucleotide sugar 6-dehydrogenase family and the kinetic and structural data we obtained for BceC, we determined Y10 as a key catalytic residue in a UGD rate-determining step, the final hydrolysis of the enzymatic thioester intermediate.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Apartado 127, P-2781-901 Oeiras, Portugal.
Organizational Affiliation: