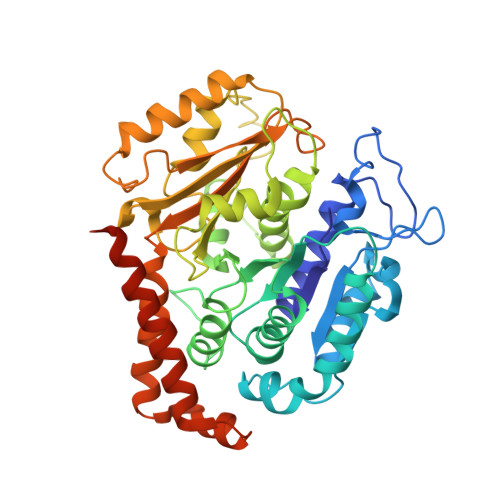
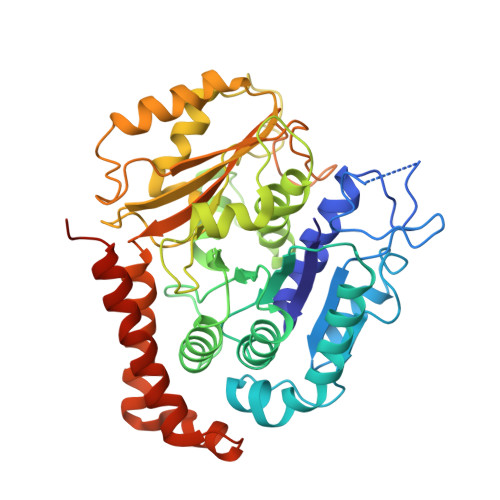
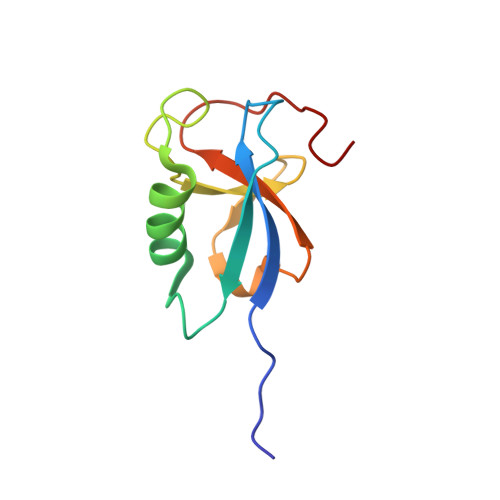
Template-Free 13-Protofilament Microtubule-Map Assembly Visualized at 8 A Resolution.
Fourniol, F.J., Sindelar, C.V., Amigues, B., Clare, D.K., Thomas, G., Perderiset, M., Francis, F., Houdusse, A., Moores, C.A.(2010) J Cell Biol 191: 463
- PubMed: 20974813 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1083/jcb.201007081
- Primary Citation Related Structures:
2XRP - PubMed Abstract:
Microtubule-associated proteins (MAPs) are essential for regulating and organizing cellular microtubules (MTs). However, our mechanistic understanding of MAP function is limited by a lack of detailed structural information. Using cryo-electron microscopy and single particle algorithms, we solved the 8 Å structure of doublecortin (DCX)-stabilized MTs. Because of DCX's unusual ability to specifically nucleate and stabilize 13-protofilament MTs, our reconstruction provides unprecedented insight into the structure of MTs with an in vivo architecture, and in the absence of a stabilizing drug. DCX specifically recognizes the corner of four tubulin dimers, a binding mode ideally suited to stabilizing both lateral and longitudinal lattice contacts. A striking consequence of this is that DCX does not bind the MT seam. DCX binding on the MT surface indirectly stabilizes conserved tubulin-tubulin lateral contacts in the MT lumen, operating independently of the nucleotide bound to tubulin. DCX's exquisite binding selectivity uncovers important insights into regulation of cellular MTs.
- Institute of Structural and Molecular Biology, Birkbeck College, London, England, UK.
Organizational Affiliation: