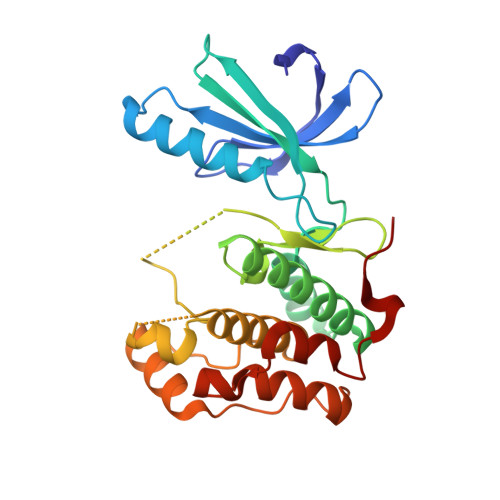
Benzimidazole Inhibitors Induce a Dfg-Out Conformation of Never in Mitosis Gene A-Related Kinase 2 (Nek2) without Binding to the Back Pocket and Reveal a Nonlinear Structure-Activity Relationship.
Solanki, S., Innocenti, P., Mas-Droux, C., Boxall, K., Barillari, C., Van Montfort, R.L., Aherne, G.W., Bayliss, R., Hoelder, S.(2011) J Med Chem 54: 1626
- PubMed: 21366329 Search on PubMed
- DOI: https://doi.org/10.1021/jm1011726
- Primary Citation Related Structures:
2XNM, 2XNN, 2XNO, 2XNP - PubMed Abstract:
We describe herein the structure-activity relationship (SAR) and cocrystal structures of a series of Nek2 inhibitors derived from the published polo-like kinase 1 (Plk1) inhibitor (R)-1. Our studies reveal a nonlinear SAR for Nek2 and our cocrystal structures show that compounds in this series bind to a DFG-out conformation of Nek2 without extending into the enlarged back pocket commonly found in this conformation. These observations were further investigated, and structure-based design led to Nek2 inhibitors derived from (R)-1 with more than a hundred-fold selectivity against Plk1.
- The Institute of Cancer Research, Cancer Research UK Cancer Therapeutics Unit, 15 Cotswold Road, Sutton, Surrey SM2 5NG, United Kingdom.
Organizational Affiliation: