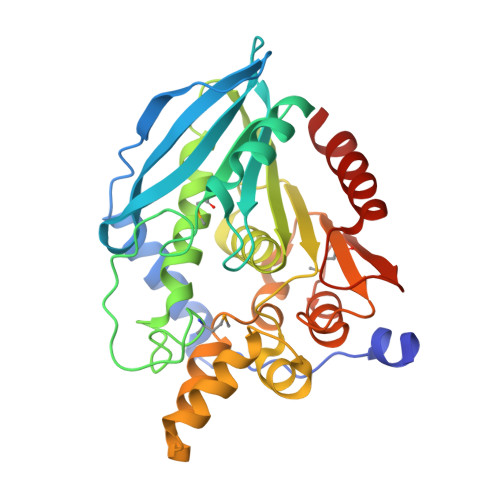
The Crystal Structure of the Cephalosporin Deacetylating Enzyme Acetyl Xylan Esterase Bound to Paraoxon Explains the Low Sensitivity of This Serine Hydrolase to Organophosphate Inactivation.
Montoro-Garcia, S., Gil-Ortiz, F., Garcia-Carmona, F., Polo, L.M., Rubio, V., Sanchez-Ferrer, A.(2011) Biochem J 436: 321
- PubMed: 21382014
- DOI: https://doi.org/10.1042/BJ20101859
- Primary Citation Related Structures:
2XLB, 2XLC - PubMed Abstract:
Organophosphorus insecticides and nerve agents irreversibly inhibit serine hydrolase superfamily enzymes. One enzyme of this superfamily, the industrially important (for β-lactam antibiotic synthesis) AXE/CAH (acetyl xylan esterase/cephalosporin acetyl hydrolase) from the biotechnologically valuable organism Bacillus pumilus, exhibits low sensitivity to the organophosphate paraoxon (diethyl-p-nitrophenyl phosphate, also called paraoxon-ethyl), reflected in a high K(i) for it (~5 mM) and in a slow formation (t(½)~1 min) of the covalent adduct of the enzyme and for DEP (E-DEP, enzyme-diethyl phosphate, i.e. enzyme-paraoxon). The crystal structure of the E-DEP complex determined at 2.7 Å resolution (1 Å=0.1 nm) reveals strain in the active Ser¹⁸¹-bound organophosphate as a likely cause for the limited paraoxon sensitivity. The strain results from active-site-size limitation imposed by bulky conserved aromatic residues that may exclude as substrates esters having acyl groups larger than acetate. Interestingly, in the doughnut-like homohexamer of the enzyme, the six active sites are confined within a central chamber formed between two 60°-staggered trimers. The exclusive access to this chamber through a hole around the three-fold axis possibly limits the size of the xylan natural substrates. The enzyme provides a rigid scaffold for catalysis, as reflected in the lack of movement associated with paraoxon adduct formation, as revealed by comparing this adduct structure with that also determined in the present study at 1.9 Å resolution for the paraoxon-free enzyme.
- Department of Biochemistry and Molecular Biology-A, Faculty of Biology, University of Murcia, Campus Espinardo, E-30100 Murcia, Spain.
Organizational Affiliation: