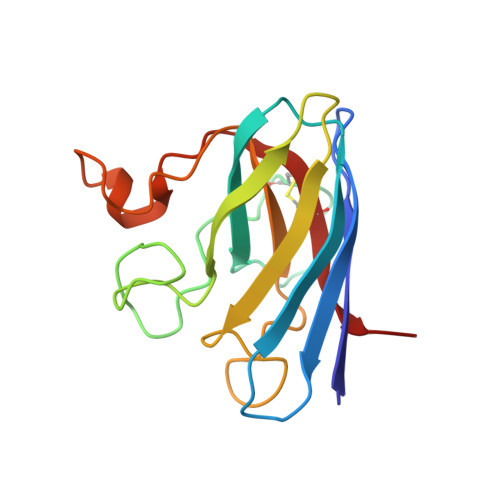
Folding Catalysis by Transient Coordination of Zn2+ to the Cu Ligands of the Als-Associated Enzyme Cu/Zn Superoxide Dismutase 1.
Leinartaite, L., Saraboji, K., Nordlund, A., Logan, D.T., Oliveberg, M.(2010) J Am Chem Soc 132: 13495
- PubMed: 20822138 Search on PubMed
- DOI: https://doi.org/10.1021/ja1057136
- Primary Citation Related Structures:
2XJK, 2XJL - PubMed Abstract:
How coordination of metal ions modulates protein structures is not only important for elucidating biological function but has also emerged as a key determinant in protein turnover and protein-misfolding diseases. In this study, we show that the coordination of Zn(2+) to the ALS-associated enzyme Cu/Zn superoxide dismutase (SOD1) is directly controlled by the protein's folding pathway. Zn(2+) first catalyzes the folding reaction by coordinating transiently to the Cu ligands of SOD1, which are all contained within the folding nucleus. Then, after the global folding transition has commenced, the Zn(2+) ion transfers to the higher affinity Zn site, which structures only very late in the folding process. Here it remains dynamically coordinated with an off rate of ∼10(-5) s(-1). This relatively rapid equilibration of metals in and out of the SOD1 structure provides a simple explanation for how the exceptionally long lifetime, >100 years, of holoSOD1 is still compatible with cellular turnover: if a dissociated Zn(2+) ion is prevented from rebinding to the SOD1 structure then the lifetime of the protein is reduced to a just a few hours.
- Department of Biochemistry and Biophysics, Arrhenius Laboratories of Natural Sciences, Stockholm University, S-106 91 Stockholm, Sweden.
Organizational Affiliation: