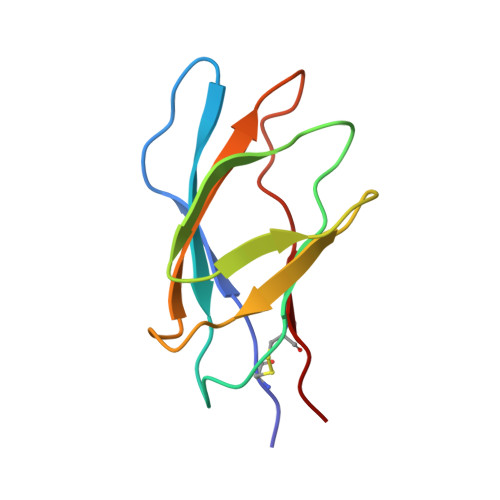
A family IIb xylan-binding domain has a similar secondary structure to a homologous family IIa cellulose-binding domain but different ligand specificity.
Simpson, P.J., Bolam, D.N., Cooper, A., Ciruela, A., Hazlewood, G.P., Gilbert, H.J., Williamson, M.P.(1999) Structure 7: 853-864
- PubMed: 10425686 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(99)80108-7
- Primary Citation Related Structures:
1XBD, 2XBD - PubMed Abstract:
Many enzymes that digest polysaccharides contain separate polysaccharide-binding domains. Structures have been previously determined for a number of cellulose-binding domains (CBDs) from cellulases. The family IIb xylan-binding domain 1 (XBD1) from Cellulomonas fimi xylanase D is shown to bind xylan but not cellulose. Its structure is similar to that of the homologous family IIa CBD from C. fimi Cex, consisting of two four-stranded beta sheets that form a twisted 'beta sandwich'. The xylan-binding site is a groove made from two tryptophan residues that stack against the faces of the sugar rings, plus several hydrogen-bonding polar residues. The biggest difference between the family IIa and IIb domains is that in the former the solvent-exposed tryptophan sidechains are coplanar, whereas in the latter they are perpendicular, forming a twisted binding site. The binding sites are therefore complementary to the secondary structures of the ligands cellulose and xylan. XBD1 and CexCBD represent a striking example of two proteins that have high sequence similarity but a different function.
- Department of Molecular Biology and Biotechnology, Krebs Institute, University of Sheffield, UK.
Organizational Affiliation: