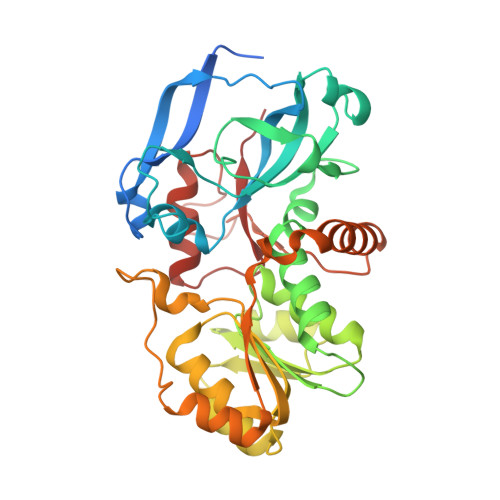
Crystal Structure of the Human Mgc45594 Gene Product in Complex with Fenoprofen
Shafqat, N., Yue, W.W., Ugochukwu, E., Niesen, F., Vollmar, M., Chaikuad, A., Pike, A.C.W., von Delft, F., Smee, C., Arrowsmith, C., Weigelt, J., Edwards, A., Bountra, C., Oppermann, U.To be published.