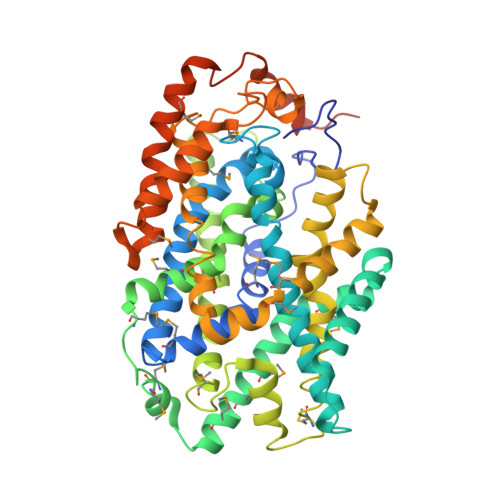
Molecular Basis of Alternating Access Membrane Transport by the Sodium-Hydantoin Transporter Mhp1.
Shimamura, T., Weyand, S., Beckstein, O., Rutherford, N.G., Hadden, J.M., Sharples, D., Sansom, M.S.P., Iwata, S., Henderson, P.J.F., Cameron, A.D.(2010) Science 328: 470
- PubMed: 20413494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1186303
- Primary Citation Related Structures:
2X79 - PubMed Abstract:
The structure of the sodium-benzylhydantoin transport protein Mhp1 from Microbacterium liquefaciens comprises a five-helix inverted repeat, which is widespread among secondary transporters. Here, we report the crystal structure of an inward-facing conformation of Mhp1 at 3.8 angstroms resolution, complementing its previously described structures in outward-facing and occluded states. From analyses of the three structures and molecular dynamics simulations, we propose a mechanism for the transport cycle in Mhp1. Switching from the outward- to the inward-facing state, to effect the inward release of sodium and benzylhydantoin, is primarily achieved by a rigid body movement of transmembrane helices 3, 4, 8, and 9 relative to the rest of the protein. This forms the basis of an alternating access mechanism applicable to many transporters of this emerging superfamily.
- Division of Molecular Biosciences, Membrane Protein Crystallography Group, Imperial College, London SW7 2AZ, UK.
Organizational Affiliation: