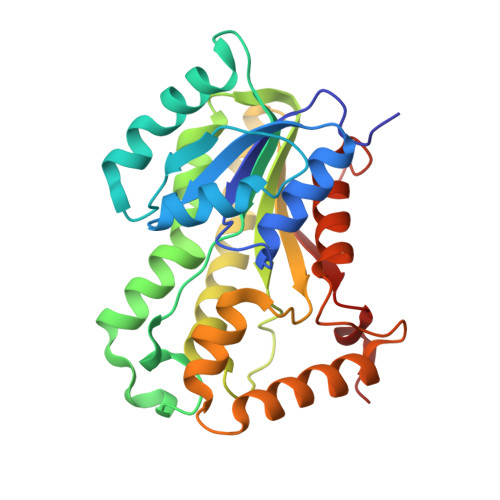
A Slow, Tight Binding Inhibitor of Inha, the Enoyl-Acyl Carrier Protein Reductase from Mycobacterium Tuberculosis.
Luckner, S.R., Liu, N., Am Ende, C.W., Tonge, P.J., Kisker, C.(2010) J Biological Chem 285: 14330
- PubMed: 20200152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.090373
- Primary Citation Related Structures:
2X22, 2X23 - PubMed Abstract:
InhA, the enoyl-ACP reductase in Mycobacterium tuberculosis is an attractive target for the development of novel drugs against tuberculosis, a disease that kills more than two million people each year. InhA is the target of the current first line drug isoniazid for the treatment of tuberculosis infections. Compounds that directly target InhA and do not require activation by the mycobacterial catalase-peroxidase KatG are promising candidates for treating infections caused by isoniazid-resistant strains. Previously we reported the synthesis of several diphenyl ethers with nanomolar affinity for InhA. However, these compounds are rapid reversible inhibitors of the enzyme, and based on the knowledge that long drug target residence times are an important factor for in vivo drug activity, we set out to generate a slow onset inhibitor of InhA using structure-based drug design. 2-(o-Tolyloxy)-5-hexylphenol (PT70) is a slow, tight binding inhibitor of InhA with a K(1) value of 22 pm. PT70 binds preferentially to the InhA x NAD(+) complex and has a residence time of 24 min on the target, which is 14,000 times longer than that of the rapid reversible inhibitor from which it is derived. The 1.8 A crystal structure of the ternary complex between InhA, NAD(+), and PT70 reveals the molecular details of enzyme-inhibitor recognition and supports the hypothesis that slow onset inhibition is coupled to ordering of an active site loop, which leads to the closure of the substrate-binding pocket.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, 97080 Würzburg, Germany.
Organizational Affiliation: