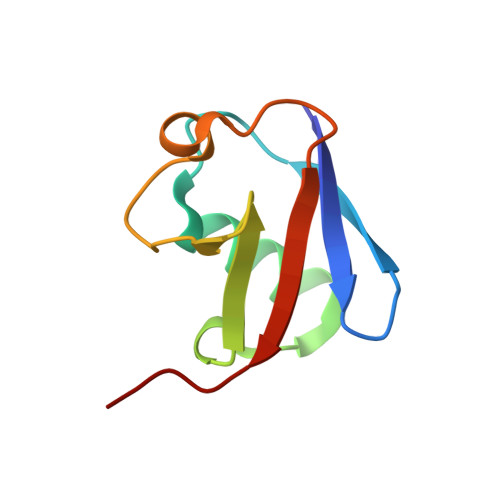
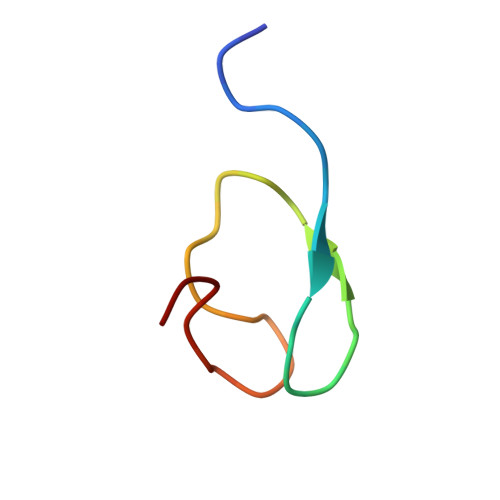
Two-Sided Ubiquitin Binding Explains Specificity of the Tab2 Nzf Domain
Kulathu, Y., Akutsu, M., Bremm, A., Hofmann, K., Komander, D.(2009) Nat Struct Mol Biol 16: 1328
- PubMed: 19935683 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1731
- Primary Citation Related Structures:
2WWZ, 2WX0, 2WX1 - PubMed Abstract:
The protein kinase TAK1 is activated by binding to Lys63 (K63)-linked ubiquitin chains through its subunit TAB2. Here we analyze crystal structures of the TAB2 NZF domain bound to Lys63-linked di- and triubiquitin, revealing that TAB2 binds adjacent ubiquitin moieties via two distinct binding sites. The conformational constraints imposed by TAB2 on a Lys63 dimer cannot be adopted by linear chains, explaining why TAK1 cannot be activated by linear ubiquitination events.
- Medical Research Council Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: