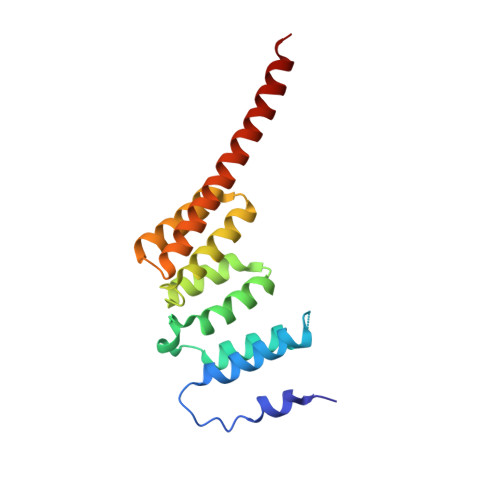
Defining the Molecular Basis of Bubr1 Kinetochore Interactions and Anaphase-Promoting Complex/Cyclosome (Apc/C)-Cdc20 Inhibition
D'Arcy, S., Davies, O.R., Blundell, T.L., Bolanos-Garcia, V.M.(2010) J Biological Chem 285: 14764
- PubMed: 20220147 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.082016
- Primary Citation Related Structures:
2WVI - PubMed Abstract:
BubR1 is essential for the mitotic checkpoint that prevents aneuploidy in cellular progeny by triggering anaphase delay in response to kinetochores incorrectly/not attached to the mitotic spindle. Here, we define the molecular architecture of the functionally significant N-terminal region of human BubR1 and present the 1.8 A crystal structure of its tetratricopeptide repeat (TPR) domain. The structure reveals divergence from the classical TPR fold and is highly similar to the TPR domain of budding yeast Bub1. Shared distinctive features include a disordered loop insertion, a 3(10)-helix, a tight turn involving glycine positive Phi angles, and noncanonical packing of and between the TPR motifs. We also define the molecular determinants of the interaction between BubR1 and kinetochore protein Blinkin. We identify a shallow groove on the concave surface of the BubR1 TPR domain that forms multiple discrete and potentially cooperative interactions with Blinkin. Finally, we present evidence for a direct interaction between BubR1 and Bub1 mediated by regions C-terminal to their TPR domains. This interaction provides a mechanism for Bub1-dependent kinetochore recruitment of BubR1. We thus present novel molecular insights into the structure of BubR1 and its interactions at the kinetochore-microtubule interface. Our studies pave the way for future structure-directed engineering aimed at dissecting the roles of kinetochore-bound and other pools of BubR1 in vivo.
- Department of Biochemistry, University of Cambridge, Cambridge CB2 1GA, United Kingdom.
Organizational Affiliation: