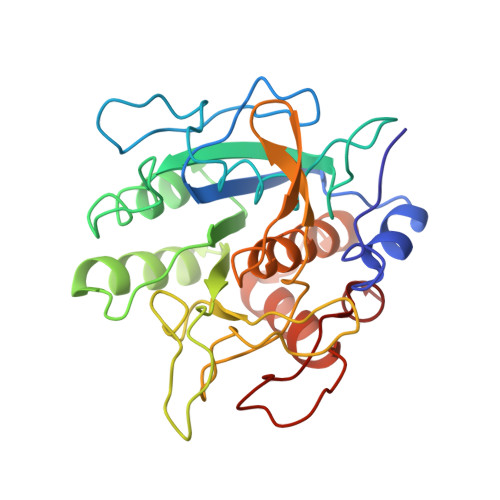
Crystallographic Analysis of Counterion Effects on Subtilisin Enzymatic Action in Acetonitrile.
Cianci, M., Tomaszewski, B., Helliwell, J.R., Halling, P.J.(2010) J Am Chem Soc 132: 2293
- PubMed: 20099851 Search on PubMed
- DOI: https://doi.org/10.1021/ja908703c
- Primary Citation Related Structures:
2WUV, 2WUW - PubMed Abstract:
When enzymes are in low dielectric nonaqueous media, it would be expected that their charged groups would be more closely associated with counterions. There is evidence that these counterions may then affect enzymatic activity. Published crystal structures of proteins in organic solvents do not show increased numbers of associated counterions, and this might reflect the difficulty of distinguishing cations like Na(+) from water molecules. In this paper, the placement of several Cs(+) and Cl(-) ions in crystals of the serine protease subtilisin Carlsberg is presented. Ions are more readily identified crystallographically through their anomalous diffraction using softer X-rays. The protein conformation is very similar to that of the enzyme without CsCl in acetonitrile, both for the previously reported ( 1SCB ) and our own newly determined model. No fewer than 11 defined sites for Cs(+) cations and 8 Cl(-) anions are identified around the protein molecule, although most of these have partial occupancy and may represent nonspecific binding sites. Two Cs(+) and two Cl(-) ions are close to the mouth of the active site cleft, where they may affect catalysis. In fact, cross-linked CsCl-treated subtilisin crystals transferred to acetonitrile show catalytic activity several fold higher than the reference crystals containing Na(+). Presoaking with another large cation, choline, also increases the enzyme activity. The active site appears only minimally sterically perturbed by the ion presence around it, so alternative activation mechanisms can be suggested: an electrostatic redistribution and/or a larger hydration sphere that enhances the protein domain.
- European Molecular Biology Laboratory, Hamburg Outstation, c/o DESY, Building 25a, Notkestrasse 85, 22603 Hamburg, Germany. m.cianci@embl-hamburg.de
Organizational Affiliation: