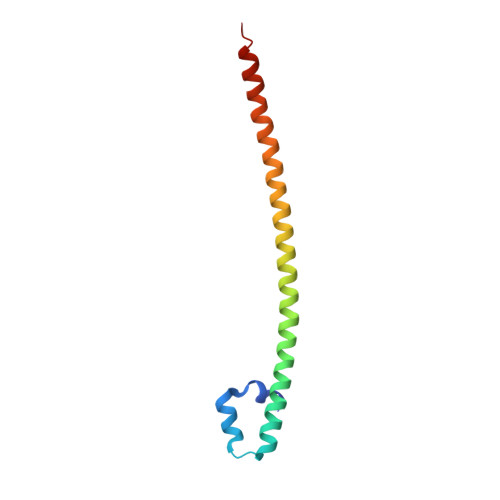
Design of a bZIP Transcription Factor with Homo/Heterodimer-Induced DNA-Binding Preference.
Pogenberg, V., Consani Textor, L., Vanhille, L., Holton, S.J., Sieweke, M.H., Wilmanns, M.(2014) Structure 22: 466
- PubMed: 24530283 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2013.12.017
- Primary Citation Related Structures:
2WT7, 2WTY - PubMed Abstract:
The ability of basic leucine zipper transcription factors for homo- or heterodimerization provides a paradigm for combinatorial control of eukaryotic gene expression. It has been unclear, however, how facultative dimerization results in alternative DNA-binding repertoires on distinct regulatory elements. To unravel the molecular basis of such coupled preferences, we determined two high-resolution structures of the transcription factor MafB as a homodimer and as a heterodimer with c-Fos bound to variants of the Maf-recognition element. The structures revealed several unexpected and dimer-specific coiled-coil-heptad interactions. Based on these findings, we have engineered two MafB mutants with opposite dimerization preferences. One of them showed a strong preference for MafB/c-Fos heterodimerization and enabled selection of heterodimer-favoring over homodimer-specific Maf-recognition element variants. Our data provide a concept for transcription factor design to selectively activate dimer-specific pathways and binding repertoires.
- EMBL Hamburg c/o DESY, Notkestraße 85, 22603 Hamburg, Germany.
Organizational Affiliation: