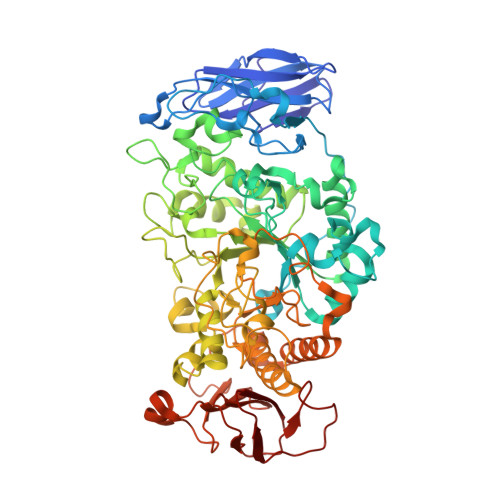
Structural Rationale for the Short Branched Substrate Specificity of the Glycogen Debranching Enzyme Glgx.
Song, H.-N., Jung, T.-Y., Park, J.-T., Park, B., Myung, P.K., Boos, W., Woo, E.-J., Park, K.-H.(2010) Proteins 78: 1847
- PubMed: 20187119 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22697
- Primary Citation Related Structures:
2WSK - PubMed Abstract:
Glycogen serves as major energy storage in most living organisms. GlgX, with its gene in the glycogen degradation operon, functions in glycogen catabolism by selectively catalyzing the debranching of polysaccharide outer chains in bacterial glycosynthesis. GlgX hydrolyzes alpha-1,6-glycosidic linkages of phosphorylase-limit dextrin containing only three or four glucose subunits produced by glycogen phosphorylase. To understand its mechanism and unique substrate specificity toward short branched alpha-polyglucans, we determined the structure of GlgX from Escherichia Coli K12 at 2.25 A resolution. The structure reveals a monomer consisting of three major domains with high structural similarity to the subunit of TreX, the oligomeric bifunctional glycogen debranching enzyme (GDE) from Sulfolobus. In the overlapping substrate binding groove, conserved residues Leu270, Asp271, and Pro208 block the cleft, yielding a shorter narrow GlgX cleft compared to that of TreX. Residues 207-213 form a unique helical conformation that is observed in both GlgX and TreX, possibly distinguishing GDEs from isoamylases and pullulanases. The structural feature observed at the substrate binding groove provides a molecular explanation for the unique substrate specificity of GlgX for G4 phosphorylase-limit dextrin and the discriminative activity of TreX and GlgX toward substrates of varying lengths.
- Medical Proteomics Research Center, Korea Institute of Bioscience and Biotechnology, Daejeon, Korea.
Organizational Affiliation: