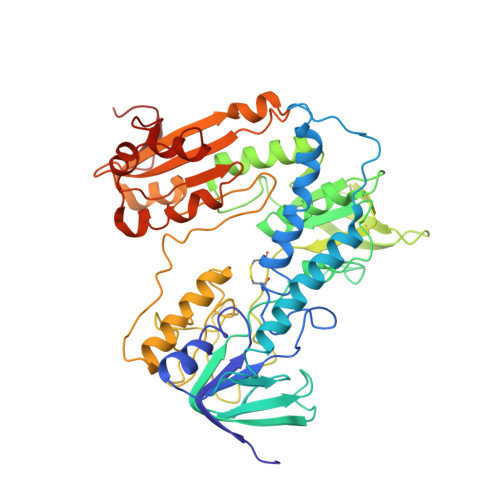
Dihydroquinazolines as a Novel Class of Trypanosoma Brucei Trypanothione Reductase Inhibitors: Discovery, Synthesis, and Characterization of Their Binding Mode by Protein Crystallography.
Patterson, S., Alphey, M.S., Jones, D.C., Shanks, E.J., Street, I.P., Frearson, J.A., Wyatt, P.G., Gilbert, I.H., Fairlamb, A.H.(2011) J Med Chem 54: 6514
- PubMed: 21851087
- DOI: https://doi.org/10.1021/jm200312v
- Primary Citation of Related Structures:
2WOI, 2WOV, 2WOW, 2WP5, 2WP6, 2WPC, 2WPE, 2WPF - PubMed Abstract:
Trypanothione reductase (TryR) is a genetically validated drug target in the parasite Trypanosoma brucei , the causative agent of human African trypanosomiasis. Here we report the discovery, synthesis, and development of a novel series of TryR inhibitors based on a 3,4-dihydroquinazoline scaffold. In addition, a high resolution crystal structure of TryR, alone and in complex with substrates and inhibitors from this series, is presented. This represents the first report of a high resolution complex between a noncovalent ligand and this enzyme. Structural studies revealed that upon ligand binding the enzyme undergoes a conformational change to create a new subpocket which is occupied by an aryl group on the ligand. Therefore, the inhibitor, in effect, creates its own small binding pocket within the otherwise large, solvent exposed active site. The TryR-ligand structure was subsequently used to guide the synthesis of inhibitors, including analogues that challenged the induced subpocket. This resulted in the development of inhibitors with improved potency against both TryR and T. brucei parasites in a whole cell assay.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee , Dow Street, Dundee DD1 5EH, U.K.
Organizational Affiliation: