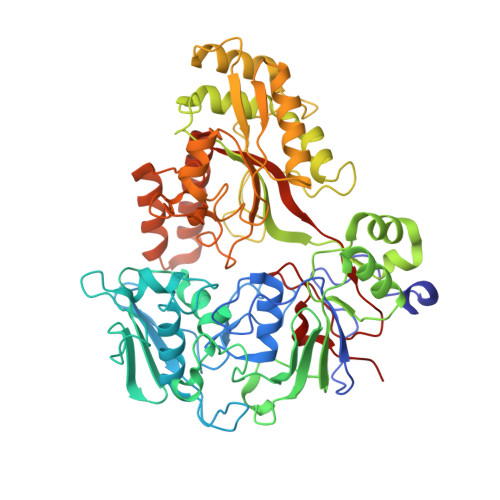
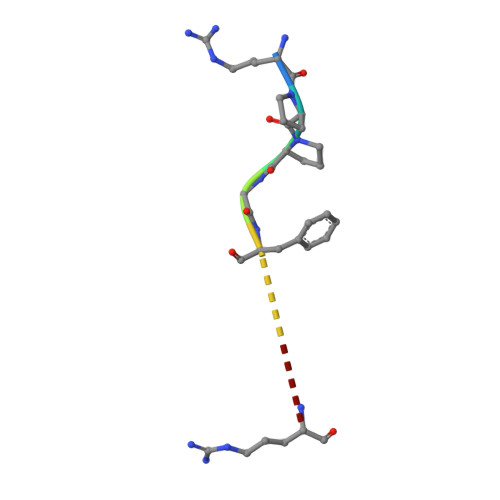
Crystal structures of an oligopeptide-binding protein from the biosynthetic pathway of the beta-lactamase inhibitor clavulanic acid.
Mackenzie, A.K., Valegard, K., Iqbal, A., Caines, M.E., Kershaw, N.J., Jensen, S.E., Schofield, C.J., Andersson, I.(2010) J Mol Biology 396: 332-344
- PubMed: 19941870 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.11.045
- Primary Citation Related Structures:
2WOK, 2WOL, 2WOP - PubMed Abstract:
Clavulanic acid (CA) is a clinically important beta-lactamase inhibitor that is produced by fermentation of Streptomyces clavuligerus. The CA biosynthesis pathway starts from arginine and glyceraldehyde-3-phosphate and proceeds via (3S,5S)-clavaminic acid, which is converted to (3R,5R)-clavaldehyde, the immediate precursor of (3R,5R)-CA. Open reading frames 7 (orf7) and 15 (orf15) of the CA biosynthesis cluster encode oligopeptide-binding proteins (OppA1 and OppA2), which are essential for CA biosynthesis. OppA1/2 are proposed to be involved in the binding and/or transport of peptides across the S. clavuligerus cell membrane. Peptide binding assays reveal that recombinant OppA1 and OppA2 bind di-/tripeptides containing arginine and certain nonapeptides including bradykinin. Crystal structures of OppA2 in its apo form and in complex with arginine or bradykinin were solved to 1.45, 1.7, and 1.7 A resolution, respectively. The overall fold of OppA2 consists of two lobes with a deep cavity in the center, as observed for other oligopeptide-binding proteins. The large cavity creates a peptide/arginine binding cleft. The crystal structures of OppA2 in complex with arginine or bradykinin reveal that the C-terminal arginine of bradykinin binds similarly to arginine. The results are discussed in terms of the possible roles of OppA1/2 in CA biosynthesis.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala, Sweden.
Organizational Affiliation: