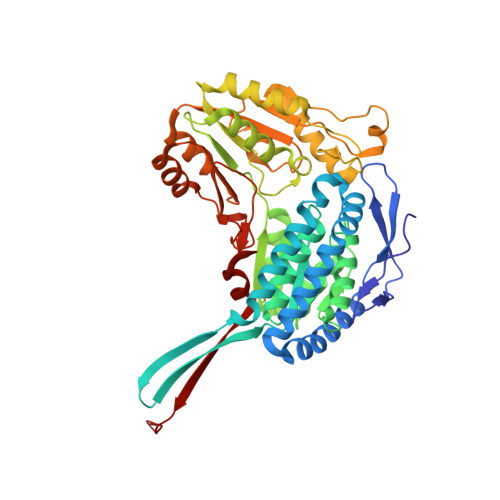
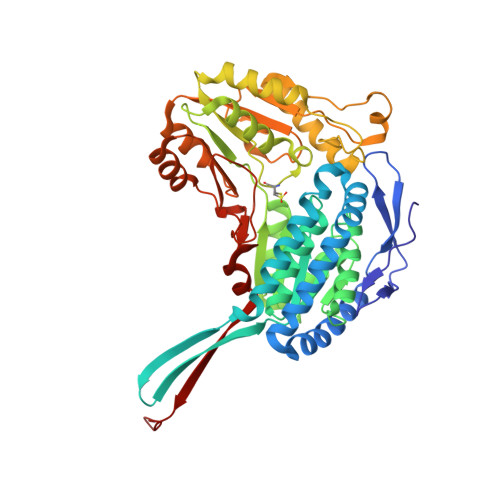
The Crystal Structure of a Ternary Complex of Betaine Aldehyde Dehydrogenase from Pseudomonas Aeruginosa Provides New Insight Into the Reaction Mechanism and Shows a Novel Binding Mode of the 2'- Phosphate of Nadp(+) and a Novel Cation Binding Site.
Gonzalez-Segura, L., Rudino-Pinera, E., Munoz-Clares, R.A., Horjales, E.(2009) J Mol Biology 385: 542
- PubMed: 19013472 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.10.082
- Primary Citation Related Structures:
2WME - PubMed Abstract:
In the human pathogen Pseudomonas aeruginosa, the NAD(P)(+)-dependent betaine aldehyde dehydrogenase (PaBADH) may play the dual role of assimilating carbon and nitrogen from choline or choline precursors--abundant at infection sites--and producing glycine betaine and NADPH, potentially protective against the high-osmolarity and oxidative stresses prevalent in the infected tissues. Disruption of the PaBADH gene negatively affects the growth of bacteria, suggesting that this enzyme could be a target for antibiotic design. PaBADH is one of the few ALDHs that efficiently use NADP(+) and one of the even fewer that require K(+) ions for stability. Crystals of PaBADH were obtained under aerobic conditions in the presence of 2-mercaptoethanol, glycerol, NADP(+) and K(+) ions. The three-dimensional structure was determined at 2.1-A resolution. The catalytic cysteine (C286, corresponding to C302 of ALDH2) is oxidized to sulfenic acid or forms a mixed disulfide with 2-mercaptoethanol. The glutamyl residue involved in the deacylation step (E252, corresponding to E268 of ALDH2) is in two conformations, suggesting a proton relay system formed by two well-conserved residues (E464 and K162, corresponding to E476 and K178, respectively, of ALDH2) that connects E252 with the bulk water. In some active sites, a bound glycerol molecule mimics the thiohemiacetal intermediate; its hydroxyl oxygen is hydrogen bonded to the nitrogen of the amide groups of the side chain of the conserved N153 (N169 of ALDH2) and those of the main chain of C286, which form the "oxyanion hole." The nicotinamide moiety of the nucleotide is not observed in the crystal, and the adenine moiety binds in the usual way. A salt bridge between E179 (E195 of ALDH2) and R40 (E53 of ALDH2) moves the carboxylate group of the former away from the 2'-phosphate of the NADP(+), thus avoiding steric clashes and/or electrostatic repulsion between the two groups. Finally, the crystal shows two K(+) binding sites per subunit. One is in an intrasubunit cavity that we found to be present in all known ALDH structures. The other--not described before for any ALDH but most likely present in most of them--is located in between the dimeric unit, helping structure a region involved in coenzyme binding and catalysis. This may explain the effects of K(+) ions on the activity and stability of PaBADH.
- Departamento de Medicina Molecular y Bioprocesos, Instituto de Biotecnología, Universidad Nacional Autónoma de México, Av. Universidad 2001, Cuernavaca, Morelos CP 62250, Mexico.
Organizational Affiliation: