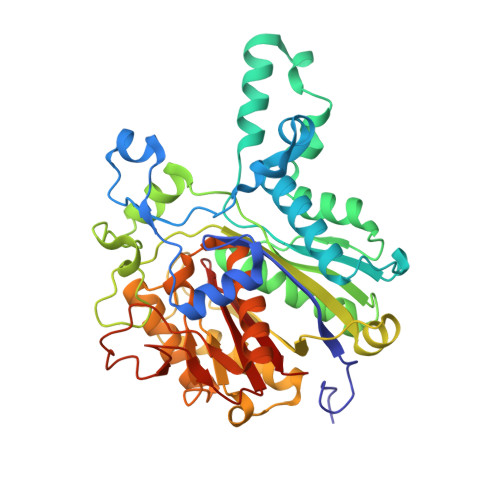
Crystal Structures of Mycobacterium Tuberculosis Kasa Show Mode of Action within Cell Wall Biosynthesis and its Inhibition by Thiolactomycin
Luckner, S.R., Machutta, C.A., Tonge, P.J., Kisker, C.(2009) Structure 17: 1004
- PubMed: 19604480 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.04.012
- Primary Citation Related Structures:
2WGD, 2WGE, 2WGF, 2WGG - PubMed Abstract:
Mycobacteria have a unique cell wall consisting of mycolic acids, very-long-chain lipids that provide protection and allow the bacteria to persist within human macrophages. Inhibition of cell wall biosynthesis is fatal for the organism and a starting point for the discovery and development of novel antibiotics. We determined the crystal structures of KasA, a key enzyme involved in the biosynthesis of long-chain fatty acids, in its apo-form and bound to the natural product inhibitor thiolactomycin. Detailed insights into the interaction of the inhibitor with KasA and the identification of a polyethylene glycol molecule that mimics a fatty acid substrate of approximately 40 carbon atoms length, represent the first atomic view of a mycobacterial enzyme involved in the synthesis of long-chain fatty acids and provide a robust platform for the development of novel thiolactomycin analogs with high affinity for KasA.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, Würzburg, Germany.
Organizational Affiliation: