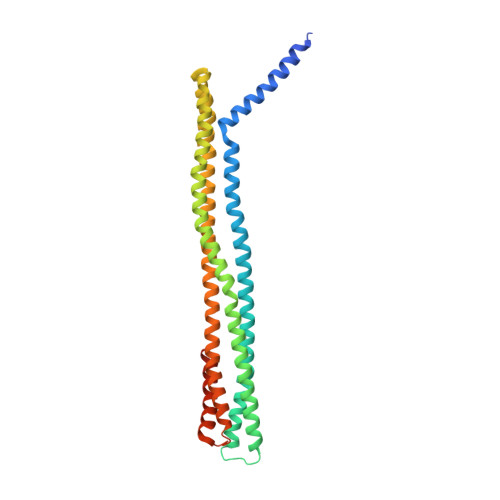
The Structure of a Cytolytic Alpha-Helical Toxin Pore Reveals its Assembly Mechanism
Mueller, M., Grauschopf, U., Maier, T., Glockshuber, R., Ban, N.(2009) Nature 459: 726
- PubMed: 19421192 Search on PubMed
- DOI: https://doi.org/10.1038/nature08026
- Primary Citation Related Structures:
2WCD - PubMed Abstract:
Pore-forming toxins (PFTs) are a class of potent virulence factors that convert from a soluble form to a membrane-integrated pore. They exhibit their toxic effect either by destruction of the membrane permeability barrier or by delivery of toxic components through the pores. Among the group of bacterial PFTs are some of the most dangerous toxins, such as diphtheria and anthrax toxin. Examples of eukaryotic PFTs are perforin and the membrane-attack complex, proteins of the immune system. PFTs can be subdivided into two classes, alpha-PFTs and beta-PFTs, depending on the suspected mode of membrane integration, either by alpha-helical or beta-sheet elements. The only high-resolution structure of a transmembrane PFT pore is available for a beta-PFT--alpha-haemolysin from Staphylococcus aureus. Cytolysin A (ClyA, also known as HlyE), an alpha-PFT, is a cytolytic -helical toxin responsible for the haemolytic phenotype of several Escherichia coli and Salmonella enterica strains. ClyA is cytotoxic towards cultured mammalian cells, induces apoptosis of macrophages and promotes tissue pervasion. Electron microscopic reconstructions demonstrated that the soluble monomer of ClyA must undergo large conformational changes to form the transmembrane pore. Here we report the 3.3 A crystal structure of the 400 kDa dodecameric transmembrane pore formed by ClyA. The tertiary structure of ClyA protomers in the pore is substantially different from that in the soluble monomer. The conversion involves more than half of all residues. It results in large rearrangements, up to 140 A, of parts of the monomer, reorganization of the hydrophobic core, and transitions of -sheets and loop regions to -helices. The large extent of interdependent conformational changes indicates a sequential mechanism for membrane insertion and pore formation.
- Institute of Molecular Biology and Biophysics, ETH Zurich, 8093 Zurich, Switzerland.
Organizational Affiliation: