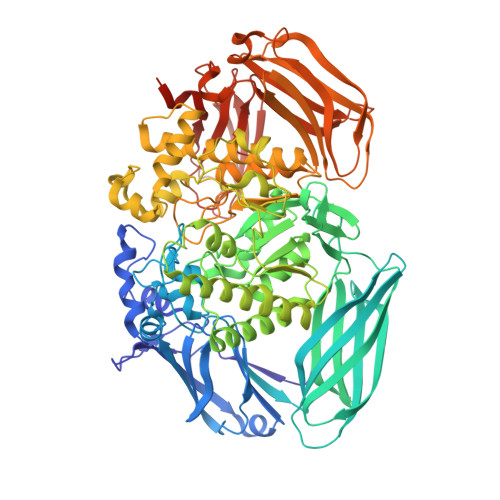
Structure of the Michaelis Complex of Beta-Mannosidase, Man2A, Provides Insight Into the Conformational Itinerary of Mannoside Hydrolysis.
Offen, W.A., Zechel, D.L., Withers, S.G., Gilbert, H.J., Davies, G.J.(2009) Cell 18: 2484
- PubMed: 19532864 Search on PubMed
- DOI: https://doi.org/10.1039/b902240f
- Primary Citation Related Structures:
2WBK - PubMed Abstract:
The Michaelis complex of the beta-mannosidase Man2A shows distortion to a (1)S(5) conformation adding to the growing body of evidence supporting catalysis through a boat conformation.
- Department of Chemistry, The University of York, Heslington, UK.
Organizational Affiliation: