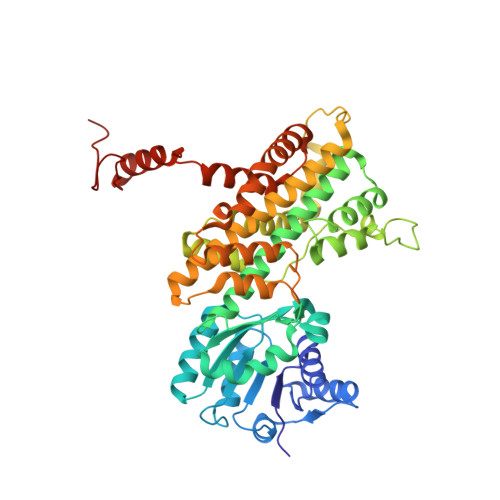
Geobacillus Stearothermophilus 6-Phosphogluconate Dehydrogenase, Complexed with 6-Phosphogluconate.
Cameron, S., Martini, V.P., Iulek, J., Hunter, W.N.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 450
- PubMed: 19407374 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109012767
- Primary Citation Related Structures:
2W8Z, 2W90 - PubMed Abstract:
Two crystal structures of recombinant Geobacillus stearothermophilus 6-phosphogluconate dehydrogenase (Gs6PDH) in complex with the substrate 6-phosphogluconate have been determined at medium resolution. Gs6PDH shares significant sequence identity and structural similarity with the enzymes from Lactococcus lactis, sheep liver and the protozoan parasite Trypanosoma brucei, for which a range of structures have previously been reported. Comparisons indicate that amino-acid sequence conservation is more pronounced in the two domains that contribute to the architecture of the active site, namely the N-terminal and C-terminal domains, compared with the central domain, which is primarily involved in the subunit-subunit associations required to form a stable dimer. The active-site residues are highly conserved, as are the interactions with the 6-phosphogluconate. There is interest in 6PDH as a potential drug target for the protozoan parasite T. brucei, the pathogen responsible for African sleeping sickness. The recombinant T. brucei enzyme has proven to be recalcitrant to enzyme-ligand studies and a surrogate protein might offer new opportunities to investigate and characterize 6PDH inhibitors. The high degree of structural similarity, efficient level of expression and straightforward crystallization conditions mean that Gs6PDH may prove to be an appropriate model system for structure-based inhibitor design targeting the enzyme from Trypanosoma species.
- University of Dundee, Scotland, UK.
Organizational Affiliation: