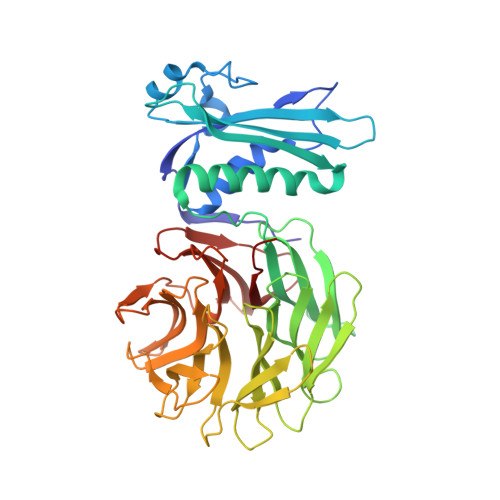
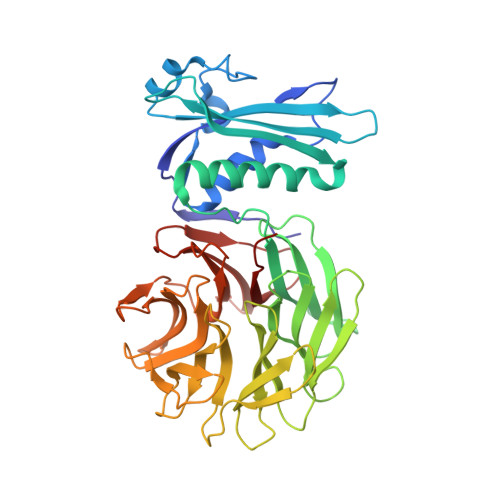
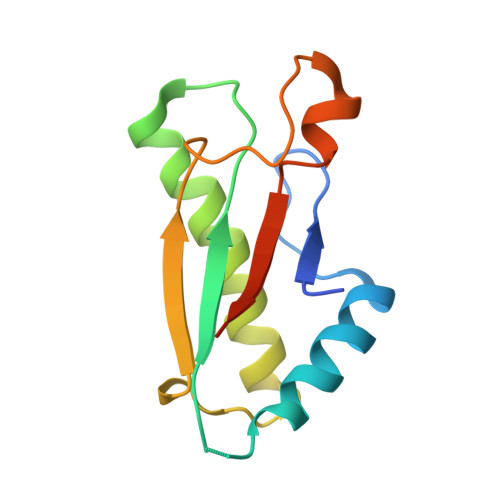
Allosteric Beta-Propeller Signalling in Tolb and its Manipulation by Translocating Colicins.
Bonsor, D.A., Hecht, O., Vankemmelbeke, M., Sharma, A., Krachler, A.M., Housden, N.G., Lilly, K.J., James, R., Moore, G.R., Kleanthous, C.(2009) EMBO J 28: 2846
- PubMed: 19696740 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2009.224
- Primary Citation Related Structures:
2W8B - PubMed Abstract:
The Tol system is a five-protein assembly parasitized by colicins and bacteriophages that helps stabilize the Gram-negative outer membrane (OM). We show that allosteric signalling through the six-bladed beta-propeller protein TolB is central to Tol function in Escherichia coli and that this is subverted by colicins such as ColE9 to initiate their OM translocation. Protein-protein interactions with the TolB beta-propeller govern two conformational states that are adopted by the distal N-terminal 12 residues of TolB that bind TolA in the inner membrane. ColE9 promotes disorder of this 'TolA box' and recruitment of TolA. In contrast to ColE9, binding of the OM lipoprotein Pal to the same site induces conformational changes that sequester the TolA box to the TolB surface in which it exhibits little or no TolA binding. Our data suggest that Pal is an OFF switch for the Tol assembly, whereas colicins promote an ON state even though mimicking Pal. Comparison of the TolB mechanism to that of vertebrate guanine nucleotide exchange factor RCC1 suggests that allosteric signalling may be more prevalent in beta-propeller proteins than currently realized.
- Department of Biology, University of York, York, UK.
Organizational Affiliation: