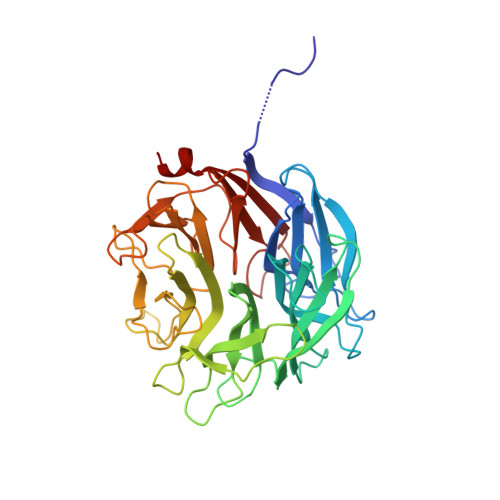
Molecular Basis of Arabinobio-Hydrolase Activity in Phytopathogenic Fungi. Crystal Structure and Catalytic Mechanism of Fusarium Graminearum Gh93 Exo-Alpha-L-Arabinanase.
Carapito, R., Imberty, A., Jeltsch, J.M., Byrns, S.C., Tam, P.H., Lowary, T.L., Varrot, A., Phalip, V.(2009) J Biological Chem 284: 12285
- PubMed: 19269961 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M900439200
- Primary Citation Related Structures:
2W5N, 2W5O - PubMed Abstract:
The phytopathogenic fungus Fusarium graminearum secretes a very diverse pool of glycoside hydrolases (GHs) aimed at degrading plant cell walls. alpha-l-Arabinanases are essential GHs participating in the complete hydrolysis of hemicellulose, a natural resource for various industrial processes, such as bioethanol or pharmaceuticals production. Arb93A, the exo-1,5-alpha-l-arabinanase of F. graminearum encoded by the gene fg03054.1, belongs to the GH93 family, for which no structural data exists. The enzyme is highly active (1065 units/mg) and displays a strict substrate specificity for linear alpha-1,5-l-arabinan. Biochemical assays and NMR experiments demonstrated that the enzyme releases alpha-1,5-l-arabinobiose from the nonreducing end of the polysaccharide. We determined the crystal structure of the native enzyme and its complex with alpha-1,5-l-arabinobiose, a degradation product of alpha-Me-1,5-l-arabinotetraose, at 1.85 and 2.05A resolution, respectively. Arb93A is a monomeric enzyme, which presents the six-bladed beta-propeller fold characteristic of sialidases of clan GHE. The configuration of the bound arabinobiose is consistent with the retaining mechanism proposed for the GH93 family. Catalytic residues were proposed from the structural analysis, and site-directed mutagenesis was used to validate their role. They are significantly different from those observed for GHE sialidases.
- UMR 7175, Ecole Supérieure de Biotechnologie de Strasbourg, Université de Strasbourg-CNRS, Boulevard Sébastien Brandt, BP 10413, 67412 Illkirch-Graffenstaden, France.
Organizational Affiliation: