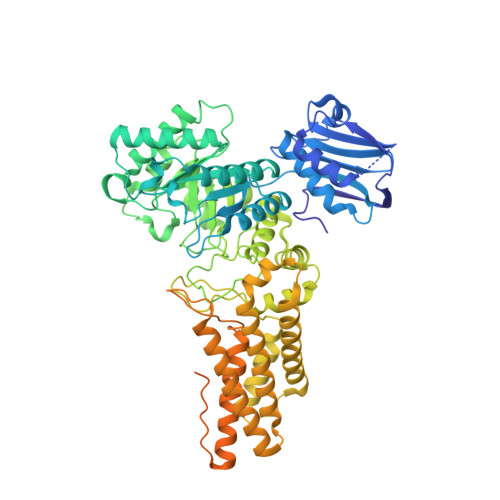
Structural Insight Into the Mechanism of Streptozotocin Inhibition of O-Glcnacase.
He, Y., Martinez-Fleites, C., Bubb, A., Gloster, T.M., Davies, G.J.(2009) Carbohydr Res 344: 627
- PubMed: 19217614 Search on PubMed
- DOI: https://doi.org/10.1016/j.carres.2008.12.007
- Primary Citation Related Structures:
2W4X - PubMed Abstract:
Despite decades of its use in diabetes research, the mechanism of cytotoxicity of streptozotocin (STZ) toward pancreatic beta-islet cells has remained a topic of discussion. Although STZ toxicity is likely a function of its capacity to promote DNA alkylation, it has been proposed that STZ induces pancreatic beta-cell death through O-GlcNAcase inhibition. In this report, we explore the binding mode of STZ to a close homolog of human O-GlcNAcase, BtGH84 from Bacteroides thetaiotaomicron. Our results show that STZ binds in the enzyme active site in its intact form, without the formation of a covalent adduct, consistent with solution studies on BtGH84 and human O-GlcNAcase, as well as with structural work on a homolog from Clostridium perfringens. The active site of the BtGH84 is considerably deformed upon STZ binding and as a result the catalytic machinery is expelled from the binding cavity.
- Department of Chemistry, Structural Biology Laboratory, The University of York, Heslington, York, YO10 5YW, UK.
Organizational Affiliation: