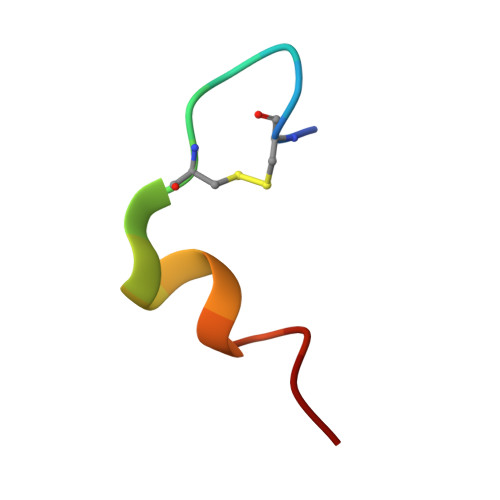
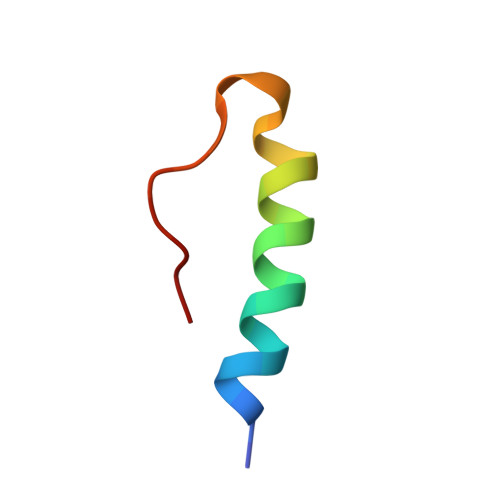
Kinetic Evidence for the Sequential Association of Insulin Binding Sites 1 and 2 to the Insulin Receptor and the Influence of Receptor Isoform.
Thorsoe, K.S., Schlein, M., Steensgaard, D.B., Brandt, J., Schluckebier, G., Naver, H.(2010) Biochemistry 49: 6234
- PubMed: 20568733 Search on PubMed
- DOI: https://doi.org/10.1021/bi1000118
- Primary Citation Related Structures:
2W44 - PubMed Abstract:
Through binding to and signaling via the insulin receptor (IR), insulin is involved in multiple effects on growth and metabolism. The current model for the insulin-IR binding process is one of a biphasic reaction. It is thought that the insulin peptide possesses two binding interfaces (sites 1 and 2), which allow it to bridge the two alpha-subunits of the insulin receptor during the biphasic binding reaction. The sequential order of the binding events involving sites 1 and 2, as well as the molecular interactions corresponding to the fast and slow binding events, is still unknown. In this study we examined the series of events that occur during the binding process with the help of three insulin analogues: insulin, an analogue mutated in site 2 (B17A insulin), and an analogue in which part of site 1 was deleted (Des A1-4 insulin), both with and without a fluorescent probe attached. The binding properties of these analogues were tested using two soluble Midi IR constructs representing the two naturally occurring isoforms of the IR, Midi IR-A and Midi IR-B. Our results showed that in the initial events leading to Midi IR-insulin complex formation, insulin site 2 binds to the IR in a very fast binding event. Subsequent to this initial fast phase, a slower rate-limiting phase occurs, consistent with a conformational change in the insulin-IR complex, which forms the final high-affinity complex. The terminal residues A1-A4 of the insulin A-chain are shown to be important for the slow binding phase, as insulin lacking these amino acids is unable to induce a conformational change of IR and has a severely impaired binding affinity. Moreover, differences in the second phase of the binding process involving insulin site 1 between the IR-A and IR-B isoforms suggest that the additional amino acids encoded by exon 11 in the IR-B isoform influence the binding process.
- Diabetes Protein Engineering, Novo Nordisk A/S, Novo Nordisk Park, 2760 Maløv, Denmark.
Organizational Affiliation: