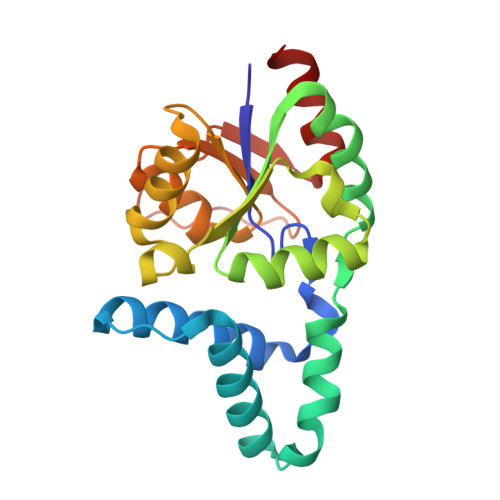
Biochemical and Structural Studies of a L-Haloacid Dehalogenase from the Thermophilic Archaeon Sulfolobus Tokodaii.
Rye, C.A., Isupov, M.N., Lebedev, A.A., Littlechild, J.A.(2009) Extremophiles 13: 179
- PubMed: 19039518 Search on PubMed
- DOI: https://doi.org/10.1007/s00792-008-0208-0
- Primary Citation Related Structures:
2W11, 2W43 - PubMed Abstract:
Haloacid dehalogenases have potential applications in the pharmaceutical and fine chemical industry as well as in the remediation of contaminated land. The L: -2-haloacid dehalogenase from the thermophilic archaeon Sulfolobus tokodaii has been cloned and over-expressed in Escherichia coli and successfully purified to homogeneity. Here we report the structure of the recombinant dehalogenase solved by molecular replacement in two different crystal forms. The enzyme is a homodimer with each monomer being composed of a core-domain of a beta-sheet bundle surrounded by alpha-helices and an alpha-helical sub-domain. This fold is similar to previously solved mesophilic L: -haloacid dehalogenase structures. The monoclinic crystal form contains a putative inhibitor L: -lactate in the active site. The enzyme displays haloacid dehalogenase activity towards carboxylic acids with the halide attached at the C2 position with the highest activity towards chloropropionic acid. The enzyme is thermostable with maximum activity at 60 degrees C and a half-life of over 1 h at 70 degrees C. The enzyme is relatively stable to solvents with 25% activity lost when incubated for 1 h in 20% v/v DMSO.
- Henry Wellcome Building for Biocatalysis, School of Biosciences, University of Exeter, Stocker Road, Exeter EX4 4QD, UK.
Organizational Affiliation: