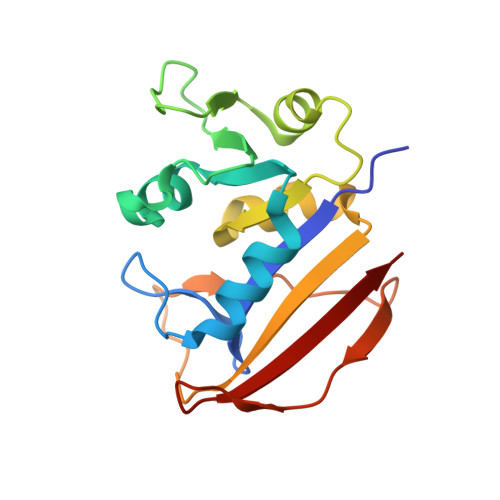
Structural Basis for Selective Inhibition of Mycobacterium Avium Dihydrofolate Reductase by a Lipophilic Antifolate
Leung, A.K.W., Ross, L.J., Zywno-Van Ginkel, S., Reynolds, R.C., Seitz, L.E., Pathak, V., Barrow, W.W., White, E.L., Suling, W.J., Piper, J.R., Borhani, D.W.To be published.