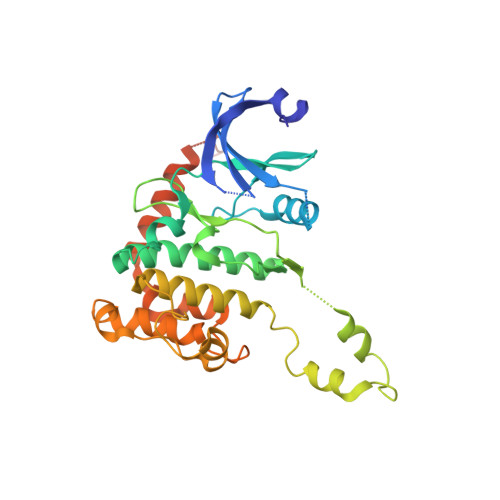
Crystal Structure of Checkpoint Kinase 2 in Complex with Nsc 109555, a Potent and Selective Inhibitor
Lountos, G.T., Tropea, J.E., Zhang, D., Jobson, A.G., Pommier, Y., Shoemaker, R.H., Waugh, D.S.(2009) Protein Sci 18: 92
- PubMed: 19177354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.16
- Primary Citation Related Structures:
2W0J - PubMed Abstract:
Checkpoint kinase 2 (Chk2), a ser/thr kinase involved in the ATM-Chk2 checkpoint pathway, is activated by genomic instability and DNA damage and results in either arrest of the cell cycle to allow DNA repair to occur or apoptosis if the DNA damage is severe. Drugs that specifically target Chk2 could be beneficial when administered in combination with current DNA-damaging agents used in cancer therapy. Recently, a novel inhibitor of Chk2, NSC 109555, was identified that exhibited high potency (IC(50) = 240 nM) and selectivity. This compound represents a new chemotype and lead for the development of novel Chk2 inhibitors that could be used as therapeutic agents for the treatment of cancer. To facilitate the discovery of new analogs of NSC 109555 with even greater potency and selectivity, we have solved the crystal structure of this inhibitor in complex with the catalytic domain of Chk2. The structure confirms that the compound is an ATP-competitive inhibitor, as the electron density clearly reveals that it occupies the ATP-binding pocket. However, the mode of inhibition differs from that of the previously studied structure of Chk2 in complex with debromohymenialdisine, a compound that inhibits both Chk1 and Chk2. A unique hydrophobic pocket in Chk2, located very close to the bound inhibitor, presents an opportunity for the rational design of compounds with higher binding affinity and greater selectivity.
- Macromolecular Crystallography Laboratory, National Cancer Institute at Frederick, P. O. Box B, Frederick, Maryland 21702-1201, USA.
Organizational Affiliation: