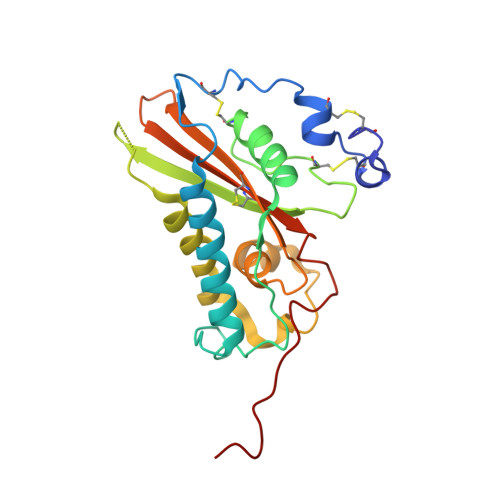
Crystal Structure of the Major Allergen from Fire Ant Venom, Sol I 3
Padavattan, S., Schmidt, M., Hoffman, D.R., Markovic-Housley, Z.(2008) J Mol Biology 383: 178
- PubMed: 18761353 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.08.023
- Primary Citation Related Structures:
2VZN - PubMed Abstract:
Fire ant venom is an extremely potent allergy-inducing agent containing four major allergens, Sol i 1 to Sol i 4, which are the most frequent cause of hypersensitivity reactions to hymenoptera in the southern USA. The crystal structure of recombinant (Baculovirus) major fire ant allergen Sol i 3 has been determined to a resolution of 3.1 A by the method of molecular replacement. The secondary-structure elements of Sol i 3 are arranged in an alpha-beta-alpha sandwich fold consisting of a central antiparallel beta-sheet surrounded on both sides by alpha helices. The overall structure is very similar to that of the homologous wasp venom allergen Ves v 5 with major differences occurring in the solvent-exposed loop regions that contain amino acid insertions. Consequently, the limited conservation of surface chemical properties and topology between Sol i 3 and Ves v 5 may explain the observed lack of relevant cross-reactivity. It is concluded that Sol i 3 recognizes immunoglobulin E antibodies with a distinct set of its own epitopes, which are different from those of Ves v 5. Indeed, the molecular area in Sol i 3 covered by non-conserved residues is large enough to accommodate four unique Sol i 3 epitopes.
- Department of Structural Biology, Biozentrum, University of Basel, CH-4056 Basel, Switzerland.
Organizational Affiliation: