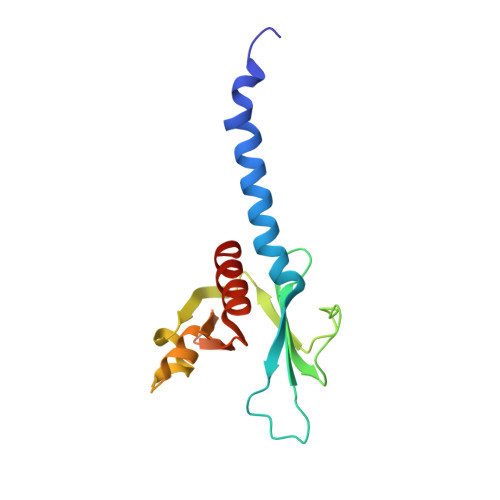
An Alpha-Helical Extension of the Elmo1 Pleckstrin Homology Domain Mediates Direct Interaction to Dock180 and is Critical in Rac Signaling.
Komander, D., Patel, M., Laurin, M., Fradet, N., Pelletier, A., Barford, D., Cote, J.-F.(2008) Mol Biol Cell 19: 4837
- PubMed: 18768751 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.e08-04-0345
- Primary Citation Related Structures:
2VSZ - PubMed Abstract:
The mammalian DOCK180 protein belongs to an evolutionarily conserved protein family, which together with ELMO proteins, is essential for activation of Rac GTPase-dependent biological processes. Here, we have analyzed the DOCK180-ELMO1 interaction, and map direct interaction interfaces to the N-terminal 200 amino acids of DOCK180, and to the C-terminal 200 amino acids of ELMO1, comprising the ELMO1 PH domain. Structural and biochemical analysis of this PH domain reveals that it is incapable of phospholipid binding, but instead structurally resembles FERM domains. Moreover, the structure revealed an N-terminal amphiphatic alpha-helix, and point mutants of invariant hydrophobic residues in this helix disrupt ELMO1-DOCK180 complex formation. A secondary interaction between ELMO1 and DOCK180 is conferred by the DOCK180 SH3 domain and proline-rich motifs at the ELMO1 C-terminus. Mutation of both DOCK180-interaction sites on ELMO1 is required to disrupt the DOCK180-ELMO1 complex. Significantly, although this does not affect DOCK180 GEF activity toward Rac in vivo, Rac signaling is impaired, implying additional roles for ELMO in mediating intracellular Rac signaling.
- Section of Structural Biology, Chester Beatty Laboratories, Institute of Cancer Research, London SW3 6JB, United Kingdom.
Organizational Affiliation: