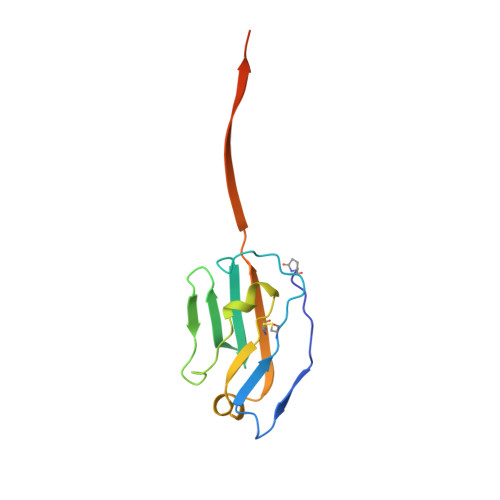
Paired Receptor Specificity Explained by Structures of Signal Regulatory Proteins Alone and Complexed with Cd47.
Hatherley, D., Graham, S.C., Turner, J., Harlos, K., Stuart, D.I., Barclay, A.N.(2008) Mol Cell 31: 266
- PubMed: 18657508 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2008.05.026
- Primary Citation Related Structures:
2JJS, 2JJT, 2JJU, 2JJV, 2JJW, 2VSC - PubMed Abstract:
CD47 is a widely distributed cell-surface protein that acts a marker of self through interactions of myeloid and neural cells. We describe the high-resolution X-ray crystallographic structures of the immunoglobulin superfamily domain of CD47 alone and in complex with the N-terminal ligand-binding domain of signal regulatory protein alpha (SIRPalpha). The unusual and convoluted interacting face of CD47, comprising the N terminus and loops at the end of the domain, intercalates with the corresponding regions in SIRPalpha. We have also determined structures of the N-terminal domains of SIRPbeta, SIRPbeta(2), and SIRPgamma; proteins that are closely related to SIRPalpha but bind CD47 with negligible or reduced affinity. These results explain the specificity of CD47 for the SIRP family of paired receptors in atomic detail. Analysis of SIRPalpha polymorphisms suggests that these, as well as the activating SIRPs, may have evolved to counteract pathogen binding to the inhibitory SIRPalpha receptor.
- Sir William Dunn School of Pathology, University of Oxford, Oxford, OX1 3RE, UK.
Organizational Affiliation: