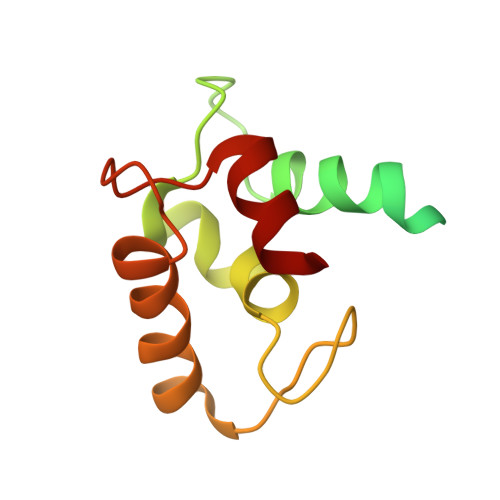
New insights into multiple coagulation factor deficiency from the solution structure of human MCFD2.
Guy, J.E., Wigren, E., Svard, M., Hard, T., Lindqvist, Y.(2008) J Mol Biology 381: 941-955
- PubMed: 18590741 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.06.042
- Primary Citation Related Structures:
2VRG - PubMed Abstract:
Human MCFD2 (multiple coagulation factor deficiency 2) is a 16-kDa protein known to participate in transport of the glycosylated human coagulation factors V and VIII along the secretory pathway. Mutations in MCFD2 or in its binding partner, the membrane-bound transporter ERGIC (endoplasmic reticulum-Golgi intermediate compartment)-53, cause a mild form of inherited hemophilia known as combined deficiency of factors V and VIII (F5F8D). While ERGIC-53 is known to be a lectin-type mannose binding protein, the role of MCFD2 in the secretory pathway is comparatively unclear. MCFD2 has been shown to bind both ERGIC-53 and the blood coagulation factors, but little is known about the binding sites or the true function of the protein. In order to facilitate understanding of the function of MCFD2 and the mechanism by which mutations in the protein cause F5F8D, we have determined the structure of human MCFD2 in solution by NMR. Our results show the folding of MCFD2 to be dependent on availability of calcium ions. The protein, which is disordered in the apo state, folds upon binding of Ca(2+) to the two EF-hand motifs of its C-terminus, while retaining some localized disorder in the N-terminus. NMR studies on two disease-causing mutant variants of MCFD2 show both to be predominantly disordered, even in the presence of calcium ions. These results provide an explanation for the previously observed calcium dependence of the MCFD2-ERGIC-53 interaction and, furthermore, clarify the means by which mutations in this protein result in inefficient secretion of blood coagulation factors V and VIII.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Stockholm, Sweden.
Organizational Affiliation: