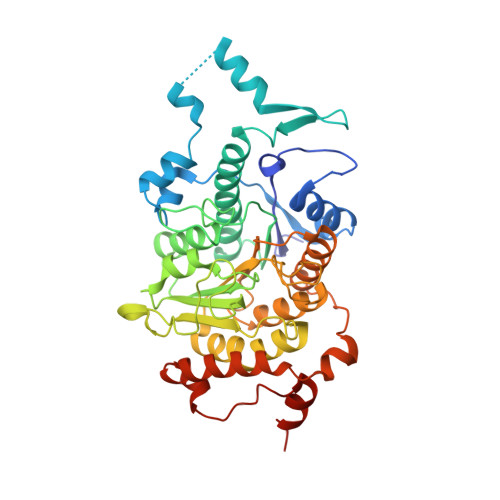
Structural and Functional Analysis of the Human Hdac4 Catalytic Domain Reveals a Regulatory Zinc-Binding Domain.
Bottomley, M.J., Lo Surdo, P., Di Giovine, P., Cirillo, A., Scarpelli, R., Ferrigno, F., Jones, P., Neddermann, P., De Francesco, R., Steinkuhler, C., Gallinari, P., Carfi, A.(2008) J Biological Chem 283: 26694
- PubMed: 18614528
- DOI: https://doi.org/10.1074/jbc.M803514200
- Primary Citation Related Structures:
2VQJ, 2VQM, 2VQO, 2VQQ, 2VQV, 2VQW - PubMed Abstract:
Histone deacetylases (HDACs) regulate chromatin status and gene expression, and their inhibition is of significant therapeutic interest. To date, no biological substrate for class IIa HDACs has been identified, and only low activity on acetylated lysines has been demonstrated. Here, we describe inhibitor-bound and inhibitor-free structures of the histone deacetylase-4 catalytic domain (HDAC4cd) and of an HDAC4cd active site mutant with enhanced enzymatic activity toward acetylated lysines. The structures presented, coupled with activity data, provide the molecular basis for the intrinsically low enzymatic activity of class IIa HDACs toward acetylated lysines and reveal active site features that may guide the design of class-specific inhibitors. In addition, these structures reveal a conformationally flexible structural zinc-binding domain conserved in all class IIa enzymes. Importantly, either the mutation of residues coordinating the structural zinc ion or the binding of a class IIa selective inhibitor prevented the association of HDAC4 with the N-CoR.HDAC3 repressor complex. Together, these data suggest a key role of the structural zinc-binding domain in the regulation of class IIa HDAC functions.
- Istituto di Ricerche di Biologia Molecolare P. Angeletti, Merck Research Laboratories, Via Pontina Km 30.600, 00040 Pomezia (Roma), Italy.
Organizational Affiliation: