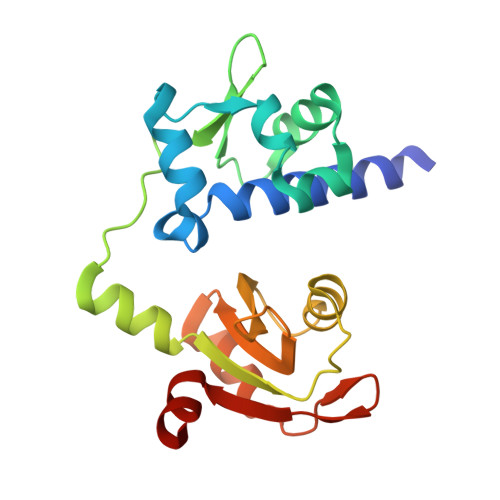
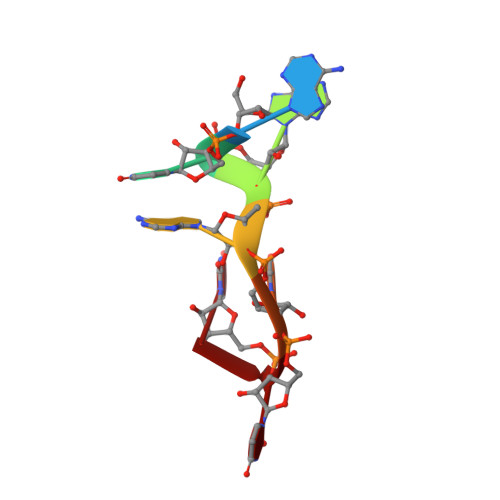
Structural Analysis Reveals Conformational Plasticity in the Recognition of RNA 3' Ends by the Human La Protein.
Kotik-Kogan, O., Valentine, E.R., Sanfelice, D., Conte, M.R., Curry, S.(2008) Structure 16: 852
- PubMed: 18547518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.02.021
- Primary Citation Related Structures:
2VOD, 2VON, 2VOO, 2VOP - PubMed Abstract:
The eukaryotic La protein recognizes the 3' poly(U) sequences of nascent RNA polymerase III transcripts to assist folding and maturation. The 3' ends of such RNAs are bound by the N-terminal domain of La (LaNTD). We have solved the crystal structures of four LaNTD:RNA complexes, each containing a different single-stranded RNA oligomer, and compared them to the structure of a previously published LaNTD:RNA complex containing partially duplex RNA. The presence of purely single-stranded RNA in the binding pocket at the interface between the La motif and RRM domains allows significantly closer contact with the 3' end of the RNA. Comparison of the different LaNTD:RNA complexes identifies a conserved set of interactions with the last two nucleotides at the 3' end of the RNA ligand that are key to binding. Strikingly, we also observe two alternative conformations of bound ssRNA, indicative of an unexpected degree of plasticity in the modes of RNA binding.
- Biophysics Section, Blackett Laboratory, Imperial College, Exhibition Road, London, United Kingdom.
Organizational Affiliation: