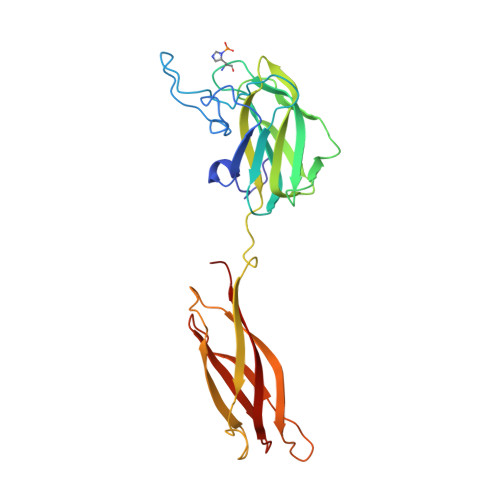
Structure Determination of Discoidin II from Dictyostelium Discoideum and Carbohydrate Binding Properties of the Lectin Domain.
Aragao, K.S., Satre, M., Imberty, A., Varrot, A.(2008) Proteins 73: 43
- PubMed: 18384150 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22038
- Primary Citation Related Structures:
2VM9, 2VMC, 2VMD, 2VME - PubMed Abstract:
The social amoeba Dictyostelium discoideum adopts a cohesive stage upon starvation and then produces Discoidin I and II, two proteins able to bind galactose and N-acetyl-galactosamine. The N-terminal domain or discoidin domain (DS) is widely distributed in eukaryotes where it plays a role in extracellular matrix binding while the C-terminal domain displays sequence similarities to invertebrate lectins. We present the first X-ray structures of the wild-type and recombinant Discoidin II in unliganded state and in complex with monosaccharides. The protein forms a homotrimer which presents two binding surfaces situated on the opposite boundaries of the structure. The binding sites of the N-terminal domain contain PEG molecules that could mimics binding of natural ligand. The C-terminal lectin domain interactions with N-acetyl-D-galactosamine and methyl-beta-galactoside are described. The carbohydrate binding sites are located at the interface between monomers. Specificity for galacto configuration can be rationalized since the axial O4 hydroxyl group is involved in several hydrogen bonds with protein side chains. Titration microcalorimetry allowed characterization of affinity and demonstrated the enthalpy-driven character of the interaction. Those results highlight the structural differentiation of the DS domain involved in many cell-adhesion processes from the lectin activity of Dictyostelium discoidins.
- CERMAV-CNRS (affiliated with University Joseph Fourier), Grenoble, France.
Organizational Affiliation: