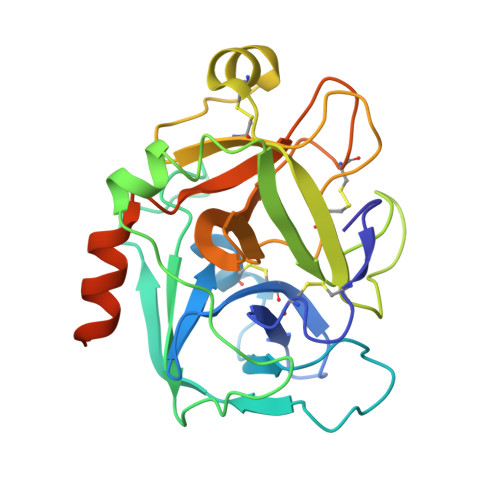
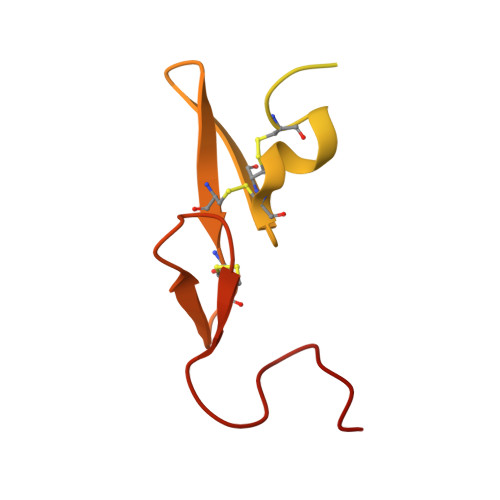
Structure and property based design of factor Xa inhibitors: biaryl pyrrolidin-2-ones incorporating basic heterocyclic motifs.
Young, R.J., Borthwick, A.D., Brown, D., Burns-Kurtis, C.L., Campbell, M., Chan, C., Charbaut, M., Convery, M.A., Diallo, H., Hortense, E., Irving, W.R., Kelly, H.A., King, N.P., Kleanthous, S., Mason, A.M., Pateman, A.J., Patikis, A.N., Pinto, I.L., Pollard, D.R., Senger, S., Shah, G.P., Toomey, J.R., Watson, N.S., Weston, H.E., Zhou, P.(2008) Bioorg Med Chem Lett 18: 28-33
- PubMed: 18053714
- DOI: https://doi.org/10.1016/j.bmcl.2007.11.019
- Primary Citation Related Structures:
2VH0 - PubMed Abstract:
Structure and property based drug design was exploited in the synthesis of sulfonamidopyrrolidin-2-one-based factor Xa (fXa) inhibitors, incorporating basic biaryl P4 groups, producing highly potent inhibitors with significant anticoagulant activities and encouraging oral pharmacokinetic profiles.
- GlaxoSmithKline, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire SG1 2NY, United Kingdom. Rob.J.Young@gsk.com
Organizational Affiliation: