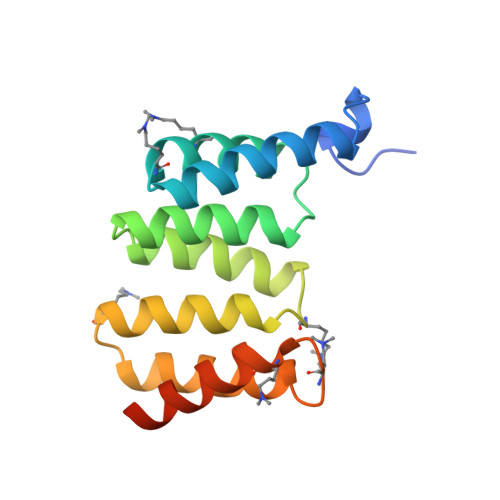
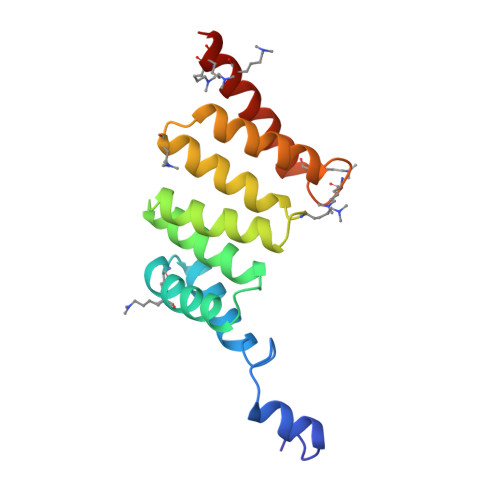
Structure of the Yersinia Enterocolitica Type III Secretion Chaperone Sycd
Buttner, C.R., Sorg, I., Cornelis, G.R., Heinz, D.W., Niemann, H.H.(2008) J Mol Biology 375: 997
- PubMed: 18054956 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.11.009
- Primary Citation Related Structures:
2VGX, 2VGY - PubMed Abstract:
Many Gram-negative bacteria use a type III secretion (T3S) system to directly inject effector molecules into eucaryotic cells in order to establish a symbiotic or pathogenic relationship with their host. The translocation of many T3S proteins requires specialized chaperones from the bacterial cytosol. SycD belongs to a class of T3S chaperones that assists the secretion of pore-forming translocators and, specifically chaperones the translocators YopB and YopD from enteropathogenic Yersinia enterocolitica. In addition, SycD is involved in the regulation of virulence factor biosynthesis and secretion. In this study, we present two crystal structures of Y. enterocolitica SycD at 1.95 and 2.6 A resolution, the first experimental structures of a T3S class II chaperone specific for translocators. The fold of SycD is entirely alpha-helical and reveals three tetratricopeptide repeat-like motifs that had been predicted from amino acid sequence. In both structures, SycD forms dimers utilizing residues from the first tetratricopeptide repeat motif. Using site-directed mutagenesis and size exclusion chromatography, we verified that SycD forms head-to-head homodimers in solution. Although in both structures, dimerization largely depends on the same residues, the two assemblies represent alternative dimers that exhibit different monomer orientations and overall shape. In these two distinct head-to-head dimers, both the concave and the convex surface of each monomer are accessible for interactions with the SycD binding partners YopB and YopD. A SycD variant carrying two point mutations in the dimerization interface is properly folded but defective in dimerization. Expression of this stable SycD monomer in Yersinia does not rescue the phenotype of a sycD null mutant, suggesting a physiological relevance of the dimerization interface.
- Division of Structural Biology, Helmholtz Centre for Infection Research, Inhoffenstrasse 7, D-38124 Braunschweig, Germany.
Organizational Affiliation: