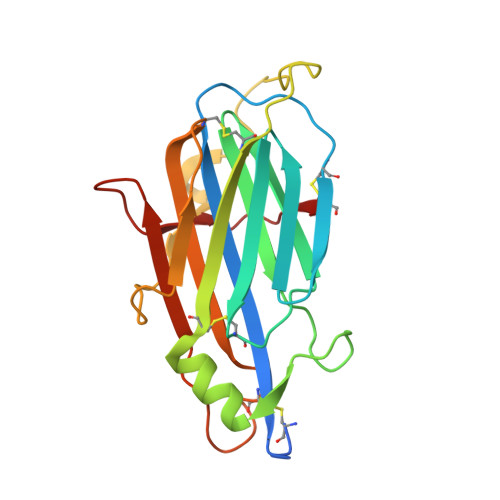
Structure and Function of A41, a Vaccinia Virus Chemokine Binding Protein.
Bahar, M.W., Kenyon, J.C., Putz, M.M., Abrescia, N.G.A., Pease, J.E., Wise, E.L., Stuart, D.I., Smith, G.L., Grimes, J.M.(2008) PLoS Pathog 4: E5
- PubMed: 18208323 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.0040005
- Primary Citation Related Structures:
2VGA - PubMed Abstract:
The vaccinia virus (VACV) A41L gene encodes a secreted 30 kDa glycoprotein that is nonessential for virus replication but affects the host response to infection. The A41 protein shares sequence similarity with another VACV protein that binds CC chemokines (called vCKBP, or viral CC chemokine inhibitor, vCCI), and strains of VACV lacking the A41L gene induced stronger CD8+ T-cell responses than control viruses expressing A41. Using surface plasmon resonance, we screened 39 human and murine chemokines and identified CCL21, CCL25, CCL26 and CCL28 as A41 ligands, with Kds of between 8 nM and 118 nM. Nonetheless, A41 was ineffective at inhibiting chemotaxis induced by these chemokines, indicating it did not block the interaction of these chemokines with their receptors. However the interaction of A41 and chemokines was inhibited in a dose-dependent manner by heparin, suggesting that A41 and heparin bind to overlapping sites on these chemokines. To better understand the mechanism of action of A41 its crystal structure was solved to 1.9 A resolution. The protein has a globular beta sandwich structure similar to that of the poxvirus vCCI family of proteins, but there are notable structural differences, particularly in surface loops and electrostatic charge distribution. Structural modelling suggests that the binding paradigm as defined for the vCCI-chemokine interaction is likely to be conserved between A41 and its chemokine partners. Additionally, sequence analysis of chemokines binding to A41 identified a signature for A41 binding. The biological and structural data suggest that A41 functions by forming moderately strong (nM) interactions with certain chemokines, sufficient to interfere with chemokine-glycosaminoglycan interactions at the cell surface (microM-nM) and thereby to destroy the chemokine concentration gradient, but not strong enough to disrupt the (pM) chemokine-chemokine receptor interactions.
- The Division of Structural Biology and The Oxford Protein Production Facility, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: