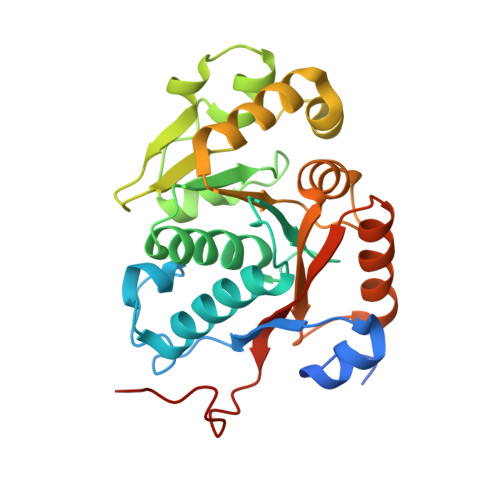
Structural Basis for the Regulation Mechanism of the Tyrosine Kinase Capb from Staphylococcus Aureus.
Olivares-Illana, V., Meyer, P., Bechet, E., Gueguen-Chaignon, V., Soulat, D., Lazereg-Riquier, S., Mijakovic, I., Deutscher, J., Cozzone, A.J., Laprevote, O., Morera, S., Grangeasse, C., Nessler, S.(2008) PLoS Biol 6: E143
- PubMed: 18547145
- DOI: https://doi.org/10.1371/journal.pbio.0060143
- Primary Citation of Related Structures:
2VED, 3BFV - PubMed Abstract:
Bacteria were thought to be devoid of tyrosine-phosphorylating enzymes. However, several tyrosine kinases without similarity to their eukaryotic counterparts have recently been identified in bacteria. They are involved in many physiological processes, but their accurate functions remain poorly understood due to slow progress in their structural characterization. They have been best characterized as copolymerases involved in the synthesis and export of extracellular polysaccharides. These compounds play critical roles in the virulence of pathogenic bacteria, and bacterial tyrosine kinases can thus be considered as potential therapeutic targets. Here, we present the crystal structures of the phosphorylated and unphosphorylated states of the tyrosine kinase CapB from the human pathogen Staphylococcus aureus together with the activator domain of its cognate transmembrane modulator CapA. This first high-resolution structure of a bacterial tyrosine kinase reveals a 230-kDa ring-shaped octamer that dissociates upon intermolecular autophosphorylation. These observations provide a molecular basis for the regulation mechanism of the bacterial tyrosine kinases and give insights into their copolymerase function.
- Laboratoire d'Enzymologie et Biochimie Structurales, CNRS, Gif sur Yvette, France.
Organizational Affiliation: