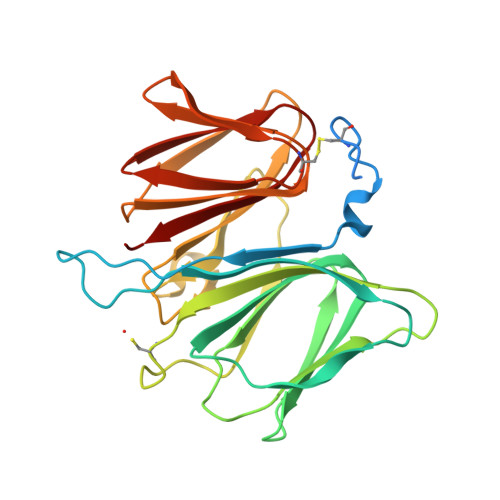
The Crystal Structure of the Protein Yhak from Escherichia Coli Reveals a New Subclass of Redox Sensitive Enterobacterial Bicupins.
Gurmu, D., Lu, J., Johnson, K.A., Nordlund, P., Holmgren, A., Erlandsen, H.(2008) Proteins 74: 18
- PubMed: 18561187 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22128
- Primary Citation Related Structures:
2VEC - PubMed Abstract:
YhaK is a protein of unknown function found in low abundance in the cytosol of Escherichia coli. DNA array studies have revealed that YhaK is strongly up-regulated by nitroso-glutathione (GSNO) and also displays a 12-fold increase in expression during biofilm growth of E. coli 83972 and VR50 in human urine. We have determined the YhaK crystal structure and demonstrated that in vitro YhaK is a good marker for monitoring oxidative stresses in E. coli. The YhaK protein structure shows a bicupin fold where the two cupin domains are crosslinked with one intramolecular disulfide bond (Cys10 to Cys204). We found that the third cysteine in YhaK, Cys122, is oxidized to a sulfenic acid. Two chloride ions are found in the structure, one close to the reactive Cys122, and the other on a hydrophobic surface close to a symmetry-related molecule. There are major structural differences at the N-terminus of YhaK compared with similar structures that also display the bicupin fold (YhhW and hPirin). YhaK showed no quercetinase and peroxidase activity. However, reduced YhaK was very sensitive to reactive oxygen species (ROS). The complete, functional E. coli glutaredoxin or thioredoxin systems protected YhaK from oxidation. E. coli thioredoxin reductase and NADPH produced ROS and caused oxidation and oligomerization of reduced YhaK. Taken together, we propose that YhaK is the first of a new sub-class of bicupins that lack the canonical cupin metal-binding residues of pirins and may be involved in chloride binding and/or sensing of oxidative stress in enterobacteria.
- Department of Biochemistry and Biophysics, The Arrhenius Laboratories for Natural Sciences, Stockholm University, 10691 Stockholm, Sweden.
Organizational Affiliation: