Discovery of Cd8+ T Cell Epitopes in Chlamydia Trachomatis Infection Through Use of Caged Class I Mhc Tetramers.
Grotenbreg, G.M., Roan, N.R., Guillen, E., Meijers, R., Wang, J.H., Bell, G.W., Starnbach, M.N., Ploegh, H.L.(2008) Proc Natl Acad Sci U S A 105: 3831
- PubMed: 18245382 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0711504105
- Primary Citation Related Structures:
2VE6 - PubMed Abstract:
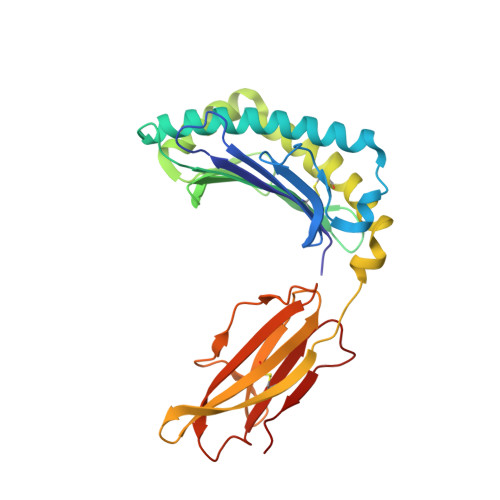
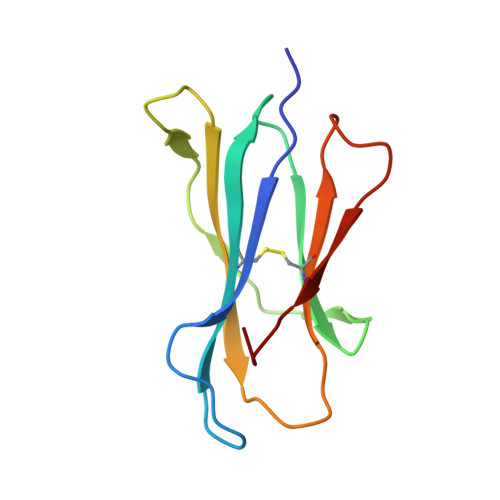
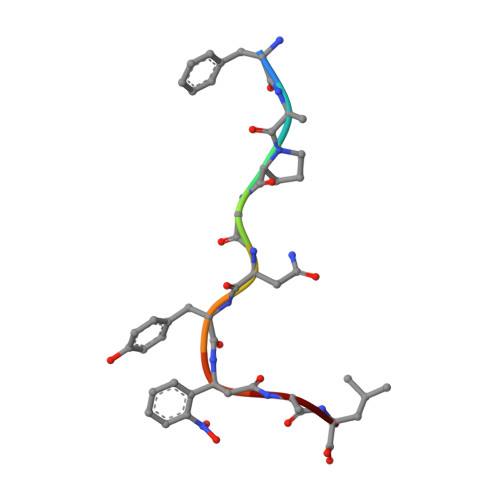
Class I MHC tetramers allow direct phenotypic identification of CD8(+) T cell populations, but their production remains laborious. A peptide exchange strategy that employs class I MHC products loaded with conditional ligands (caged MHC molecules) provides a fast and straightforward method to obtain diverse arrays of class I MHC tetramers and facilitates CD8(+) T cell epitope discovery. Here, we describe the development of photocleavable analogs of the FAPGNYPAL (SV9) epitope that bind H-2K(b) and H-2D(b) with full retention of their structural and functional integrity. We ranked all possible H-2K(b) octameric and H-2D(b) nonameric epitopes that span the genome of Chlamydia trachomatis and prepared MHC tetramers from approximately 2,000 of the highest scoring peptides by replacement of the SV9 analog with the peptide of choice. The resulting 2,000-member class I MHC tetramer array allowed the discovery of two variants of an epitope derived from polymorphic membrane protein I (PmpI) and an assessment of the kinetics of emergence and the effector function of the corresponding CD8(+) T cells.
- Whitehead Institute for Biomedical Research, 9 Cambridge Center, Cambridge, MA 02139, USA.
Organizational Affiliation: