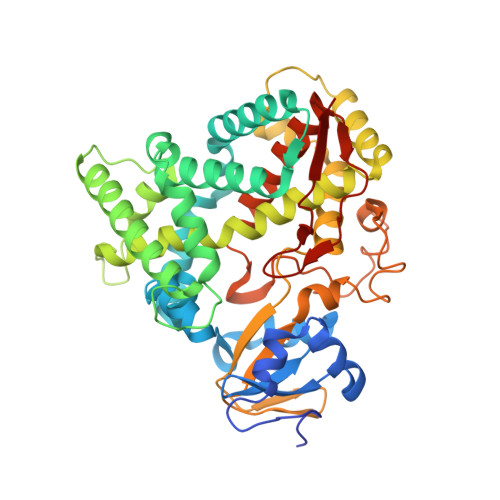
Crystal Structures of Substrate-Free and Retinoic Acid-Bound Cyanobacterial Cytochrome P450 Cyp120A1.
Kuhnel, K., Ke, N., Cryle, M.J., Sligar, S.G., Schuler, M.A., Schlichting, I.(2008) Biochemistry 47: 6552
- PubMed: 18512957
- DOI: https://doi.org/10.1021/bi800328s
- Primary Citation Related Structures:
2VE3, 2VE4 - PubMed Abstract:
The crystal structures of substrate-free and all-trans-retinoic acid-bound CYP120A1 from Synechocystis sp. PCC 6803 were determined at 2.4 and 2.1 A resolution, respectively, representing the first structural characterization of a cyanobacterial P450. Features of CYP120A1 not observed in other P450 structures include an aromatic ladder flanking the channel leading to the active site and a triple-glycine motif within SRS5. Using spectroscopic methods, CYP120A1 is shown to bind 13-cis-retinoic acid, 9-cis-retinoic acid, and retinal with high affinity and dissociation constants of less than 1 microM. Metabolism of retinoic acid by CYP120A1 suggests that CYP120A1 hydroxylates a variety of retinoid derivatives in vivo. On the basis of the retinoic acid-bound CYP120A1 crystal structure, we propose that either carbon 2 or the methyl groups (C16 or C17) of the beta-ionone ring are modified by CYP120A1.
- Department of Biomolecular Mechanisms, Max Planck Institute for Medical Research, Jahnstrasse 29, 69120 Heidelberg, Germany.
Organizational Affiliation: