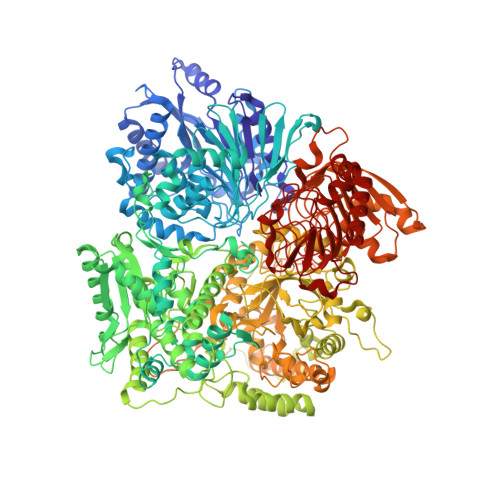
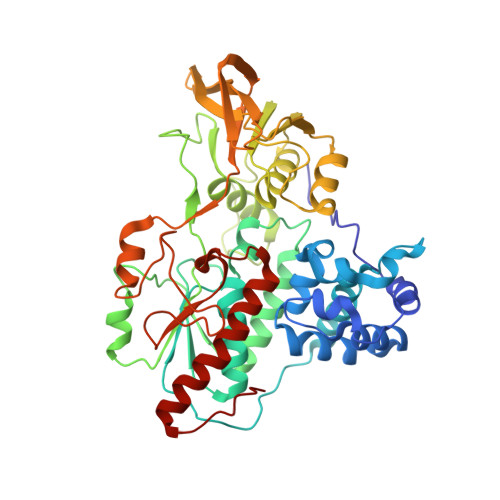
The Subnanometer Resolution Structure of the Glutamate Synthase 1.2-Mda Hexamer by Cryoelectron Microscopy and its Oligomerization Behavior in Solution: Functional Implications.
Cottevieille, M., Larquet, E., Jonic, S., Petoukhov, M.V., Caprini, G., Paravisi, S., Svergun, D.I., Vanoni, M.A., Boisset, N.(2008) J Biological Chem 283: 8237-8249
- PubMed: 18199747 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M708529200
- Primary Citation Related Structures:
2VDC - PubMed Abstract:
The three-dimensional structure of the hexameric (alphabeta)(6) 1.2-MDa complex formed by glutamate synthase has been determined at subnanometric resolution by combining cryoelectron microscopy, small angle x-ray scattering, and molecular modeling, providing for the first time a molecular model of this complex iron-sulfur flavoprotein. In the hexameric species, interprotomeric alpha-alpha and alpha-beta contacts are mediated by the C-terminal domain of the alpha subunit, which is based on a beta helical fold so far unique to glutamate synthases. The alphabeta protomer extracted from the hexameric model is fully consistent with it being the minimal catalytically active form of the enzyme. The structure clarifies the electron transfer pathway from the FAD cofactor on the beta subunit, to the FMN on the alpha subunit, through the low potential [4Fe-4S](1+/2+) centers on the beta subunit and the [3Fe-4S](0/1+) cluster on the alpha subunit. The (alphabeta)(6) hexamer exhibits a concentration-dependent equilibrium with alphabeta monomers and (alphabeta)(2) dimers, in solution, the hexamer being destabilized by high ionic strength and, to a lower extent, by the reaction product NADP(+). Hexamerization seems to decrease the catalytic efficiency of the alphabeta protomer only 3-fold by increasing the K(m) values measured for l-Gln and 2-OG. However, it cannot be ruled out that the (alphabeta)(6) hexamer acts as a scaffold for the assembly of multienzymatic complexes of nitrogen metabolism or that it provides a means to regulate the activity of the enzyme through an as yet unknown ligand.
- Département de Biologie Structurale, IMPMC-UMR 7590, CNRS, Universités Paris 6 et Paris 7, IPGP, Paris, France.
Organizational Affiliation: