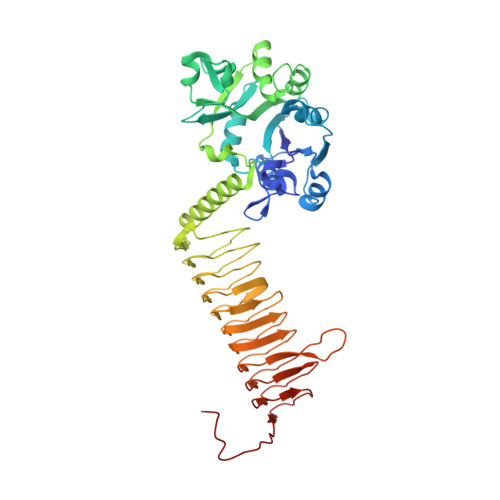
Structure of a Small-Molecule Inhibitor Complexed with Glmu from Haemophilus Influenzae Reveals an Allosteric Binding Site.
Mochalkin, I., Lightle, S., Narasimhan, L., Bornemeier, D., Melnick, M., Vanderroest, S., Mcdowell, L.(2008) Protein Sci 17: 577
- PubMed: 18218712 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.073271408
- Primary Citation Related Structures:
2VD4 - PubMed Abstract:
N-Acetylglucosamine-1-phosphate uridyltransferase (GlmU) is an essential enzyme in aminosugars metabolism and an attractive target for antibiotic drug discovery. GlmU catalyzes the formation of uridine-diphospho-N-acetylglucosamine (UDP-GlcNAc), an important precursor in the peptidoglycan and lipopolisaccharide biosynthesis in both Gram-negative and Gram-positive bacteria. Here we disclose a 1.9 A resolution crystal structure of a synthetic small-molecule inhibitor of GlmU from Haemophilus influenzae (hiGlmU). The compound was identified through a high-throughput screening (HTS) configured to detect inhibitors that target the uridyltransferase active site of hiGlmU. The original HTS hit exhibited a modest micromolar potency (IC(50) approximately 18 microM in a racemic mixture) against hiGlmU and no activity against Staphylococcus aureus GlmU (saGlmU). The determined crystal structure indicated that the inhibitor occupies an allosteric site adjacent to the GlcNAc-1-P substrate-binding region. Analysis of the mechanistic model of the uridyltransferase reaction suggests that the binding of this allosteric inhibitor prevents structural rearrangements that are required for the enzymatic reaction, thus providing a basis for structure-guided design of a new class of mechanism-based inhibitors of GlmU.
- Pfizer Inc., Michigan Laboratories, Ann Arbor, Michigan 48105, USA. Igor.Mochalkin@pfizer.com
Organizational Affiliation: