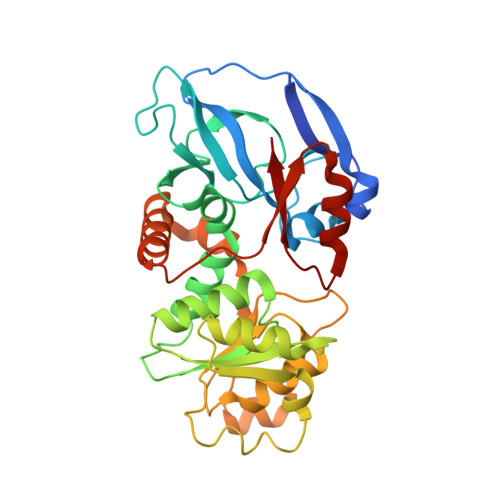
Structural Enzymological Studies of 2-Enoyl Thioester Reductase of the Human Mitochondrial Fas II Pathway: New Insights Into its Substrate Recognition Properties.
Chen, Z.J., Pudas, R., Sharma, S., Smart, O.S., Juffer, A.H., Hiltunen, J.K., Wierenga, R.K., Haapalainen, A.M.(2008) J Mol Biology 379: 830
- PubMed: 18479707 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.04.041
- Primary Citation Related Structures:
2VCY - PubMed Abstract:
Structural and kinetic properties of the human 2-enoyl thioester reductase [mitochondrial enoyl-coenzyme A reductase (MECR)/ETR1] of the mitochondrial fatty acid synthesis (FAS) II pathway have been determined. The crystal structure of this dimeric enzyme (at 2.4 A resolution) suggests that the binding site for the recognition helix of the acyl carrier protein is in a groove between the two adjacent monomers. This groove is connected via the pantetheine binding cleft to the active site. The modeled mode of NADPH binding, using molecular dynamics calculations, suggests that Tyr94 and Trp311 are critical for catalysis, which is supported by enzyme kinetic data. A deep, water-filled pocket, shaped by hydrophobic and polar residues and extending away from the catalytic site, was recognized. This pocket can accommodate a fatty acyl tail of up to 16 carbons. Mutagenesis of the residues near the end of this pocket confirms the importance of this region for the binding of substrate molecules with long fatty acyl tails. Furthermore, the kinetic analysis of the wild-type MECR/ETR1 shows a bimodal distribution of catalytic efficiencies, in agreement with the notion that two major products are generated by the mitochondrial FAS II pathway.
- Biocenter Oulu and Department of Biochemistry, University of Oulu, P.O. Box 3000, FI-90014, Oulu, Finland.
Organizational Affiliation: