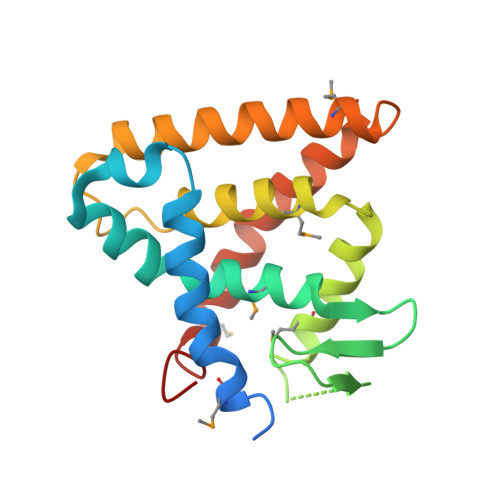
Structural Insight Into the Constitutive Repression Function of the Nuclear Receptor Rev-Erbbeta
Woo, E.-J., Jeong, D.G., Lim, M.-Y., Kim, S.J., Kim, K.-J., Yoon, S.-M., Park, B.-C., Ryu, S.E.(2007) J Mol Biology 373: 735
- PubMed: 17870090 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.08.037
- Primary Citation Related Structures:
2V0V, 2V7C - PubMed Abstract:
The Rev-erb family is an orphan nuclear receptor acting as a negative regulator of transcription. Rev-erbalpha and Rev-erbbeta are crucial components of the circadian clock and involved in various lipid homeostasis. They are unique nuclear receptors that lack the activation function 2 helix (AF2-helix) required for ligand-dependent activation by other members of nuclear receptors. Here, we report the crystal structure of Rev-erbbeta (NR1D2) in a dimeric arrangement. The putative ligand-binding pocket (LBP) of Rev-erbbeta is filled with bulky hydrophobic residues resulting in a residual cavity size that is too small to allow binding of any known ligand molecules. However, an alternative conformation of the putative LBP observed in another crystal form suggests the flexibility of this region. The kinked conformation of helix H11 allows helix H11 to bend toward helix H3 over the putative ligand binding pocket by filling and closing the cavity with its side-chains. In the absence of the AF2-helix and a cognate ligand, Rev-erbbeta appears to stabilize the hydrophobic cluster in the putative ligand binding pocket and provide a structural platform for co-repressor binding by adopting the unique geometry of helix H11, a suitable conformation for the constitutive repression activity.
- Translational Research Center, Korea Research Institute of Bioscience and Biotechnology (KRIBB), 52 Eoeun-dong, Yuseonggu, Daejeon, Korea. ejwoo@kribb.re.kr
Organizational Affiliation: