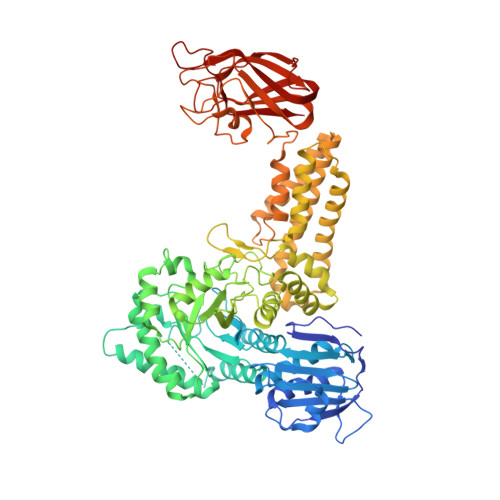
Portrait of an Enzyme: A Complete Structural Analysis of a Multi-Modular Beta-N-Acetylglucosaminidase from Clostridium Perfringens
Ficko-Blean, E., Gregg, K.J., Adams, J.J., Hehemann, J.H., Smith, S.J., Czjzek, M., Boraston, A.B.(2009) J Biological Chem 284: 9876
- PubMed: 19193644 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M808954200
- Primary Citation Related Structures:
2V5C, 2V5D, 2W1N - PubMed Abstract:
Common features of the extracellular carbohydrate-active virulence factors involved in host-pathogen interactions are their large sizes and modular complexities. This has made them recalcitrant to structural analysis, and therefore our understanding of the significance of modularity in these important proteins is lagging. Clostridium perfringens is a prevalent human pathogen that harbors a wide array of large, extracellular carbohydrate-active enzymes and is an excellent and relevant model system to approach this problem. Here we describe the complete structure of C. perfringens GH84C (NagJ), a 1001-amino acid multimodular homolog of the C. perfringens micro-toxin, which was determined using a combination of small angle x-ray scattering and x-ray crystallography. The resulting structure reveals unprecedented insight into how catalysis, carbohydrate-specific adherence, and the formation of molecular complexes with other enzymes via an ultra-tight protein-protein interaction are spatially coordinated in an enzyme involved in a host-pathogen interaction.
- Department of Biochemistry & Microbiology, University of Victoria, Victoria, British Columbia V8W 3P6, Canada.
Organizational Affiliation: